Medicine Group. 2024 September 27;5(9):1200-1213. doi: 10.37871/jbres2008.
Populations of Triple Negative and Hormone Receptor Positive HER2 Negative Breast Tumors Share Immune Gene Profiles
Frank Schmitz1,2, Wilbert B Copeland2, Justine Dell Aringa2, Kathryn Newhall2, Mary L Disis3 and Sasha Elizabeth Stanton3,4*
2Celgene Corporation, 400 Dexter Ave N #1200, Seattle, WA 98109, USA
3University of Washington, Cancer Vaccine Institute, 850 Republican Street Box 358050, Seattle, WA 98109, USA
4Earle A. Chiles Research Institute, 4805 NE Glisan St 2N103F Portland, OR 97213
- Breast cancer
- Triple negative breast cancer
- Hormone receptor positive HER2
- breast cancer; Immune infiltrate
Abstract
Background: Immune therapy has revolutionized the treatment of certain cancers including melanoma and lung cancer. Breast cancer has fewer tumors with high immune infiltrate and Immune Checkpoint Inhibitor (ICI) therapy has shown the most significant responses in the Triple Negative (TN) subtype although roles are emerging in the other subtypes. The presence of Tumor Infiltrating Lymphocytes (TIL) is predictive of response to ICI therapy in metastatic TN and associated with better survival in local TN disease. Hormone Receptor (HR) positive HER2 negative (HR+HER2-) breast tumors do not show improved prognosis with increasing TIL and only a small portion of HR+HER2- tumors respond to ICI in both metastatic and local disease. We therefore sought to identify genetic differences between TN and HR+HER2- tumors to identify how these may contribute to their differences in immune infiltrate.
Methods: We contrasted immune-associated gene expression between 119 TN and 475 HR+HER2- breast tumors from The Cancer Genome Atlas (TCGA) and confirmed our findings from 299 TN and 1369 HR+HER2- breast tumors in METABRIC.
Results: We found that TN and HR+HER2- tumors can be grouped into immune-high or -low tumors, with both subtypes represented in the immune-high group (40% HR+HER2- tumors of both Luminal A and Luminal B type and 71% TN tumors) in the TCGA. The genes upregulated in the immune-high cluster included GZMB, CXCL9, and IDO1 (Log2FC > 2; Adj. p < 10X10-40). The largest expression difference between the immune-high TN and HR+HER2- tumors was that TN tumors had more abundant Th1 and Th2 CD4+ T cells while HR+HER2- tumors had more abundant fibroblasts (Log2FC > 0.3; Adj p < 10X10-10).
Conclusions: Our data suggest that an immune-high signature is not dictated by breast cancer subtype. However, fibroblasts, particularly fibroblast subsets associated with worse outcome, are more abundant in the immune-high HR+HER2- tumors.
Abbreviations
TIL: Tumor Infiltrating Lymphocytes; HR+, HER2-: Hormone Receptor; HER2+: HER2 Positive; TN: Triple Negative; ICI: Immune Checkpoint Inhibitor; FC: Fold Change; GSVA: Gene Set Variation Analysis; CAF: Cancer Associated Fibroblast
Introduction
The histological subtyping of breast cancer, based on Hormone Receptors (HR) (estrogen receptor, ER, progesterone receptor, PR) and human epidermal growth factor receptor ERBB2 (HER2) is essential to clinical management, determining prognosis, and treatment of breast cancer [1-4]. The histologic subtypes have differences in the type and amount of immune infiltrate but a spectrum of immune infiltrate exist in all breast cancer subtypes [5]. The benefit of Immune Checkpoint Inhibitors (ICI) have been seen in metastatic cancers including renal cell, melanoma, and non-small-cell lung cancer [5-7]. The predictive biomarkers associated with response to ICI include PD-L1 expression, IFNγ expression, and increased somatic tumor mutations [6,7]. Also, a determinant for response to ICI therapy is the presence of an adaptive immune response within a tumor prior to treatment [8-11]. Breast cancer does not have the same level of adaptive immune infiltrate seen in melanoma and non-small cell lung cancer although the level of immune infiltrate in breast cancer is associated with clinical prognosis [12]. Breast tumors with greater than 50% Tumor Infiltrating Lymphocytes (TIL) show improved pathologic complete response to neoadjuvant chemotherapy (p ≤ 0.01)(11) and increased intratumoral CD8+ T cells predict improved Overall Survival (OS) (p ≤ 0.01) [13]. In the adjuvant setting, each 10% increase of TIL predicted a 15% improved Disease Free Survival (DFS) (p = 0.025) and 17% improved OS (p = 0.023) in Triple Negative (TN) but not hormone receptor positive HER2 negative (HR+HER2-) breast cancer [14,15].
In clinical studies, response to ICIs have best been seen in metastatic TN breast cancer which had a 19% response rate to pembrolizumab as a monotherapy [16]. In contrast, metastatic HR+HER2 breast cancer either with pembrolizumab as a single agent (KEYNOTE-028) or in combination with a CDK4/6 inhibitor had an overall response rate of ~12% [17,18]. In neoadjuvant therapy, pembrolizumab has improved pathologic Complete Response (pCR) in both HR+HER2- and TN. However, while the addition of pembrolizumab improved pCR in HR+HER2- tumors from 15.6% to 24.3%, the addition of pembrolizumab improves pCR in TN from 51.2% to 64.8% [19,20]. The same magnitude of benefit of ICI therapy seen in TN breast cancer was not seen in HR+HER2- breast cancer, leading to the classification of HR+HER2- breast cancer as “cold” tumors that do not respond to immune therapy.
Targeted sequencing technology has proved useful for elucidating the complex relationships between cancer and the immune system through RNA expression. We used the HTG Oncology and Immune-associated gene sets because they had previously been used to detect gene expression patterns that predicted responses in oncology clinical trials. For example, HTG Oncology’s immune-associated gene set was used to both demonstrate ICI therapy responders and predict resistance to ICIs in clinical trials across a wide variety of cancers including urothelial, colon, and lung [21,22] and has been used to identify neoadjuvant ICI response in TN breast cancer in the GeparNeuvo trial [23]. The HTG oncology panel has also been used in breast cancer clinical trials to determine changes in expression that predict treatment effect with the CDK4/6 therapy palbociclib in HR+HER2- breast cancer in PALOMA2 and PALOMA3 [24].
We questioned whether we could identify distinct immune gene expression patterns in HR+HER2- and TN breast cancers that could account for the differences in immune infiltrate between these subtypes. Analyzing gene expression data from 594 breast cancer samples using the HTG Oncology and Immuno-oncology panels, 119 TN and 475 HR+HER2- from The Cancer Genome Atlas (TCGA), we found two distinct subgroups of immune-high and immune-low tumors in both subtypes. This was confirmed in 1668 HER2- tumors in the METABRIC dataset, 299 TN and 1369 HR+HER2- breast tumors. After estimating relative cellular phenotype abundances from bulk RNA sequencing, we found cancer-associated fibroblasts more abundant in HR+HER2- tumors, particularly higher in immune-high HR+HER2- tumors than immune-high TN tumors, while helper T cell populations were more abundant in immune-high TN tumors. These differences between immune high TN and HR+HER2- tumors may contribute to the differences in response to ICI but further work is needed to determine if CAF may provide targets to improve HR+HER2- response to ICI.
Materials and Methods
Collection of datasets from The Cancer Genome Atlas (TCGA)
Normalized, bulk RNA sequencing expression data and clinical annotations were downloaded from the UCSC Xena TCGA portal for 119 TN and 475 HR+HER2- (412 ER+PR+, 63 ER+PR-, 0 ER-PR+) breast cancer patients. Therefore the TCGA analysis set included 20% TN and 80% HR+HER2- tumors. HR+HER2- and TN subtypes were defined using the clinical guidelines from the American Society of Clinical Oncology and are publicly available from previous studies of breast tumors in TCGA [25]. Mutation data (MAF) files were downloaded from the GDC Data Portal and mutational burden was analyzed using maftools [26]. The METABRIC gene expression and annotation was downloaded from the cBioPortal for Cancer Genomics and included 1369 HR+HER2- and 299 TN tumors. Annotations for immune subtypes (C1-C6) were collected from Thorsson V, et al. [27]. Annotations for the fibroblast subtypes were collected from Bartoschek M, et al. [28].
Oncology- and immune-associated gene sets
RNA expression was reported for 20,530 genes. We divided genes into two categories: oncology- and immune-associated (Supplementary Table 1(A,B)). 2,500 were captured in the HTG Edge Seq Oncology Biomarker and 543 genes were captured in the HTG Immune assays (HTG Molecular Diagnostics, Inc., Tucson, AZ).
| Table 1: RNA sequencing cell types abundance estimation. | |
| Class | Cell type |
| Dendritic | aDC |
| cDC | |
| DC | |
| iDC | |
| pDC | |
| B cells | B cells |
| Class-switched memory B cells | |
| Plasma cells | |
| proB cells | |
| Neutophils | |
| NK cells | NK |
| NKT | |
| CD4+ T cells | CD4+ T cells |
| CD4+ naïve T cells | |
| CD4+ memory T cells | |
| CD4+ effector memory T cells | |
| CD4+ central memory T cells | |
| CD8+ T cells | CD8+ T cells |
| CD8+ naïve T cells | |
| CD8+ effector memory T cells | |
| CD8+ central memory T cells | |
| Th1 cells | |
| Th2 cells | |
| Treg cells | |
| Tgd cells | |
| M1 macrophages | |
| M2 macrophages | |
| Monocytes | |
K-means clustering and silhouette analysis
To determine the optimal number of clusters for k-means, silhouette analysis (from the R ‘cluster’ and ‘factoextra’ packages) was performed on the onco- and immune-associated genes independently. K-means clustering was performed (k = 2; Hartigan and Wong algorithm, ‘kmeans’ from the R ‘stats’ package) on RNA expression values for onco- and immune-associated genes groups separately. Using Silhouette analysis identified k = 2 as optimal number of clusters using the HTG immune genes and oncogene sets (Supplementary figure 1(A-D)). Hierarchical clustering of the most variable genes in the immune-associated gene set reveals two immune-associated gene expression patterns.
Gene Set Variation Analysis (GSVA) and cell type frequency estimation
GSVA scores were calculated to measure gene set variation relative to the variation observed for all genes. To calculate sample-wise enrichment of a gene set, we used ‘GSVA’ in R [29]. Cell type enrichment was estimated with the ‘xCell’ package in R [30]. We restricted our analysis to immune relevant cell types (Table 1). To confirm xCell enrichment scores, we calculated GSVA scores for independent gene sets for CD8+ T cells (CD8A, CD8B, CD3D) or Treg (IL2RA, CCR4, CTLA4, CD28, ICOS, FOXP3, CD5).
Differential gene expression and cell enrichment
To identify differentially expressed genes or differentially abundant cell types between two groups we performed similar tests. We used limma in R [31] to calculate the log2fold change (log2FC) which describes the differences in gene expression or cell-type frequency between the designated groups. P-values were corrected for multiple hypothesis testing using the “BH” (Benjamini Hochberg, controlling the false discovery rate) method.
Results
Immune-associated gene expression is not driven by clinical subtype
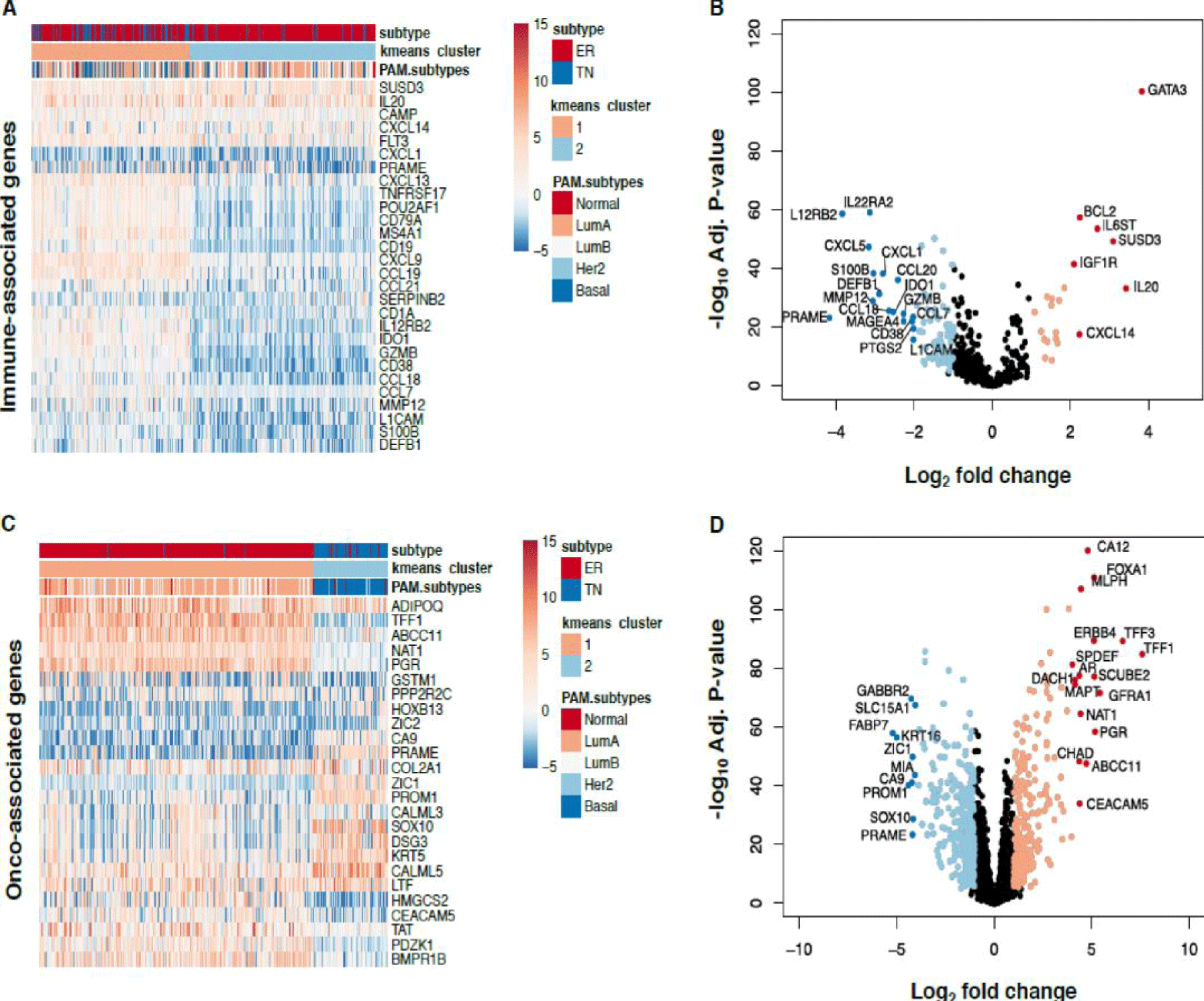
To investigate RNA expression profiles of breast cancer samples, we analyzed 119 TN and 475 HR+HER2- breast cancer samples and limited our analysis to the genes included in HTG Oncology’s immune and oncology gene sets. For the immune-associated genes, we began by performing k-means clustering to determine the optimal number of groups to immunologically characterize breast tumors (k = 2; see Material and Methods and Supplementary figure 1(A,B)). Cluster 1 had higher median log2 expression of 543 immune genes than Cluster 2 (0.29 vs. -0.32), and thus Cluster 1 was referred to as ‘immune-high’ and Cluster 2 as ‘immune-low’ (Figure 1A). Cluster 1 contains 276 breast tumors, 84 TN tumors and 192 HR+HER2- tumors (30% of the immune-high tumors are TN and 70% of the immune-high tumors are HR+HER2-). By PAM-50 typing, both cluster 1 and cluster 2 HR+HER2- tumors included luminal A and luminal B tumors and not a predominance of luminal B in cluster 1 (Figure 1A). Cluster 2 contains 318 breast tumors, 35 TN and 283 HR+HER2. When we evaluated which immune-associated genes were differentially expressed between HR+HER2- and TN breast tumors within the immune-high cluster, GATA3 (logFC = 3.8; adj. p = 4x10-101) was most expressed in HR+HER2- tumors (Figure 1B). GATA3 has previously been associated with hormone receptor expression and lower tumor grade [32].
We evaluated the same TN and HR+HER2- tumors using the HTG oncology-associated genes to show these tumors genetically matched the breast cancer subtypes found by immunohistochemistry. We performed k-means clustering of 2,500 oncology-associated genes and found that, as expected, there were two major clusters: Cluster 1 is 97% HR+HER2- tumors (461 tumors) and Cluster 2 is 88% TN tumors (107 tumors) (Figure 1C). The oncology-associated genes overexpressed more in HR+HER2- are hormone related including ESR1, ERBB4, and PGR which were more expressed in cluster 1 (log FC > 5, p < 0.001 for each) (Figure 1D).
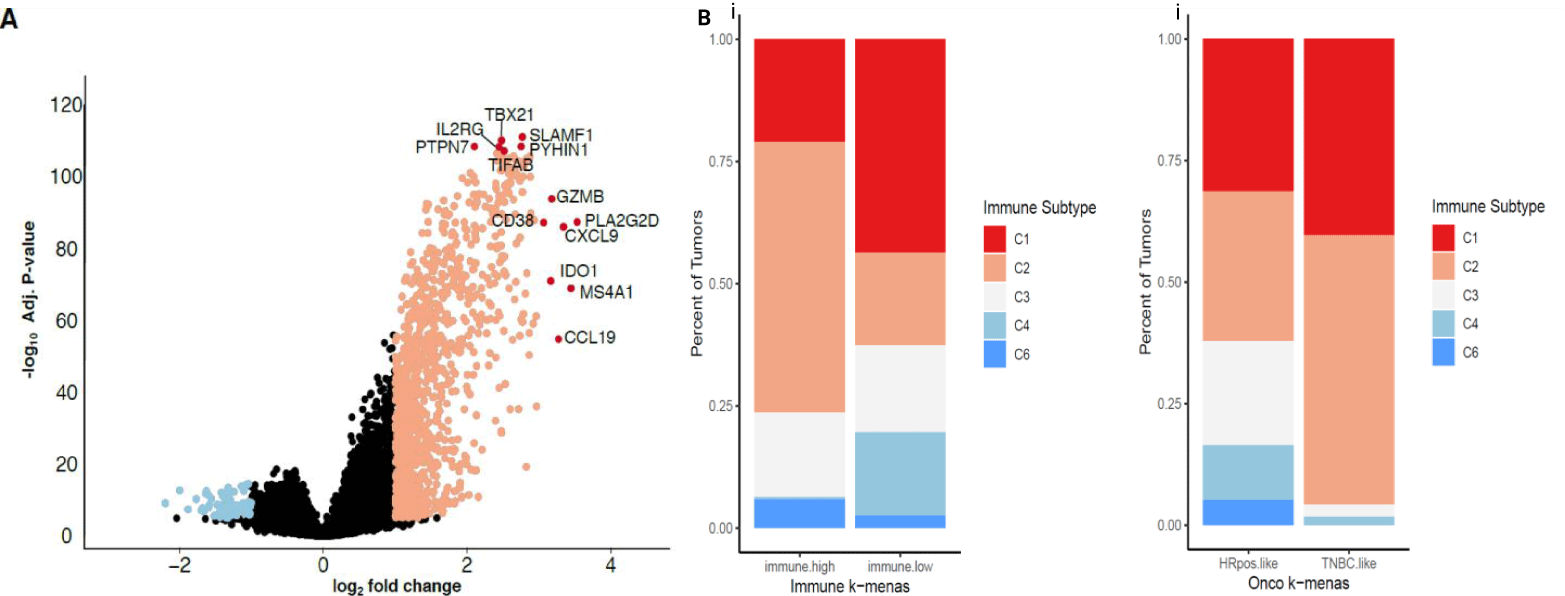
We also performed Principal Component Analysis (PCA) on the oncology and immune-associated gene sets individually. For the oncology-associated genes, the principal component capturing the greatest variance (PC1: 16.4%) between tumor gene expression strongly separates TN and HR+ tumors (Supplementary figure 2A). This is illustrated by a high area under the curve (AUC) of 0.96 (Supplementary figure 2B). However, in a PCA with immune-associated genes, we find the first principal component (PC1: 32%) is less associated with breast cancer subtype than the second principal component (PC2: 8%; AUCs = 0.74 vs. 0.91; Supplementary figure 2(C,D)). Thus, the histologic subtype is not the primary driver of immune-associated gene expression. When evaluating all genes available in TCGA (n = 20,530) the gene expression differences between immune-high and low breast tumors included immune response genes including T cell signaling (e.g., IL2RG) and function (e.g., GZMB) (Figure 2A).
To provide independent validation of the immune-high and immune-low clusters, we evaluated k-means clusters in the METABRIC dataset. The METABRIC dataset includes 1658 HER2- breast tumors: 299 TN and 1369 HR+HER2-. This dataset similarly showed two immune clusters (see Materials and Methods); an immune-high cluster 1 (240 TN and 384 HR+HER2- tumors) and an immune-low cluster 2 (59 TN and 985 HR+HER2- tumors). In this data set, 80% of TN tumors and 28% of HR+HER2- tumors were in the immune-high cluster (Supplementary figure 3A). Like the TCGA data set, when evaluating expression of the oncology-associated gene set, breast tumors generally clustered with their HR+HER2- or TN subtype similar to results by immunohistochemistry. There were 97% HR+HER2- and 12% TN in cluster 1 and 87% TN and 3% HR+HER2- in cluster 2 (Supplementary figure 3B).
Using immune clusters previously defined in 33 cancer types in TCGA [27], we evaluated if these clusters distributed both within the oncology and immune-k-means clusters (Figure 2B). We find the immune-high samples are enriched in C2 as compared to immune-low samples (55% vs 19%; figure 2Bi). Immune-high contains 21% of C1 (wound healing) where immune-low contains 44% of C1. The immune subset C5 was absent in the TCGA dataset, as seen in previous studies. When comparing oncology-gene based clustering, TN tumors are enriched for IFNγ dominant (C2) tumors compared to HR+HER2- (Figure 2Bii). This is illustrated by the difference in the proportion of C2 tumors between HR+HER2- (C2 = 31%) and TN (C2 = 55%) oncology k-means clusters. This is concordant with previous reports describing a stronger T cell presence in TN versus HR+HER2- [12]. However, we find the cluster associated with the best overall survival (C3) is almost exclusively found in HR+HER2- tumors, with 21% of the HR+HER2- tumors having a C3 immune environment compared to 2.5% of the TN tumors. Forty-six percent of the C3 tumors are immune-high and, as only six of the C3 tumors were TN, the majority of immune-high C3 tumors are HR+HER2- (n = 43, 91% of all immune-high C3 tumors) [27].
Inferred cell abundance differences between TN and HR+HER2- tumors within the immune-high cluster and the immune-low cluster.
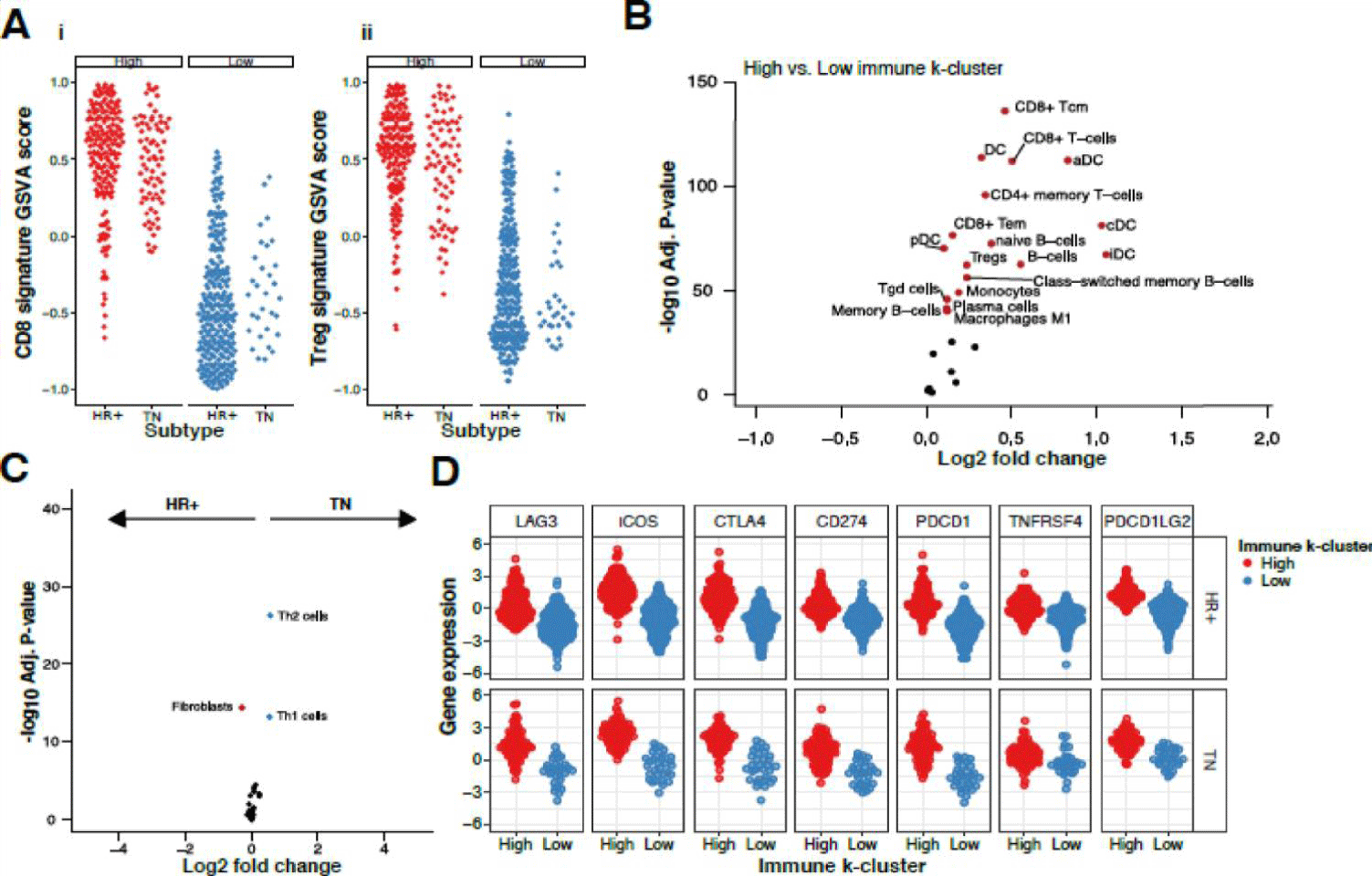
To better understand the underlying differences between TN and HR+HER2- immune-high and -low clusters, we used gene signatures for CD8+ (Figure 3Ai) and CD4+ Treg (Figure 3Aii) T cells as proxy for immune cell type abundance in the tumor. As expected, GSVA scores for CD8+ T and CD4+ Treg cell signatures in the immune-high cluster were twice that of the immune-low cluster. To estimate immune cell enrichment with cell type signatures, we used xCell(30) and confirmed that xCell scores were behaving appropriately in this dataset including T cells, B cells, dendritic cells, and macrophages that were significantly higher in the immune-high cluster (raw p < 10-50; figure 3B).
Next, we sought to characterize the immune differences between TN and HR+HER2- within the patients of the immune-high cluster. Contrasting HR+HER2- and TN in the immune-high cluster, we found TH1 and TH2 CD4+ T cells are more abundant in TN than HR+HER2- and fibroblasts are more abundant in HR+HER2- tumors than TN tumors (log2FC > 0.3; p < 10-10; figure 3C). There was no association between Th1 and Th2 infiltrate in either immune-high (ρ = 0.15; p = 0.1859) or immune-low (ρ = 0.07; p = 0.6735) TN breast cancer. However, in HR+HER2- tumors there was a positive association with increased TH1 tumors also having increased TH2 infiltrate in both immune-high (ρ = 0.29; p <= 0.0001) and immune-low (ρ = 0.21; p = 0.0005) (Supplementary figure 4). In a similar analysis of the immune-low cluster, we observe more Th1 and Th2 CD4+ T cells in TN than in HR+HER2- tumors and more fibroblasts in HR+HER2- tumors than TN tumors (Supplementary figure 5). This suggests the immune infiltrate may be impacted by the differences in fibroblasts between HR+HER2- and TN subtype regardless of the tumors being in the immune-high or immune-low groups (Supplementary tables (2,3)).
Immune checkpoint genes are differentially expressed between TN and HR+HER2- in the immune-high cluster but not in the immune-low cluster.
Immune checkpoint genes LAG3, ICOS, CTLA4, PD1 (PDCD1), PDL1 (CD274), PDL2 (PDCD1LG2), and OX40 (TNFRSF4), have been shown to predict responsiveness of tumors to ICI therapy [33,34]. Testing for differential expression levels of these genes, we observed statistically significant higher expression in the immune-high group than the immune-low group as expected (Figure 3D, Welch’s 2 sided T-test; adj. p < 9X10-36). In the immune low group the immune checkpoint target genes LAG3, ICOS, PD-1, and PD-L1 were not statistically different between TN and HR+HER2- tumors (adjusted p > 0.05) but CTLA4 and OX40 were higher in TNBC than HR+HER2- (adjusted P-value p = 0.016 each). Irrespective of hormone receptor status, immune checkpoint expression was significantly higher in the immune high cluster than in the immune low cluster (adjusted P-value p < 0.001) and in the immune high group there were no significant differences between HR+HER2- and TN tumors.
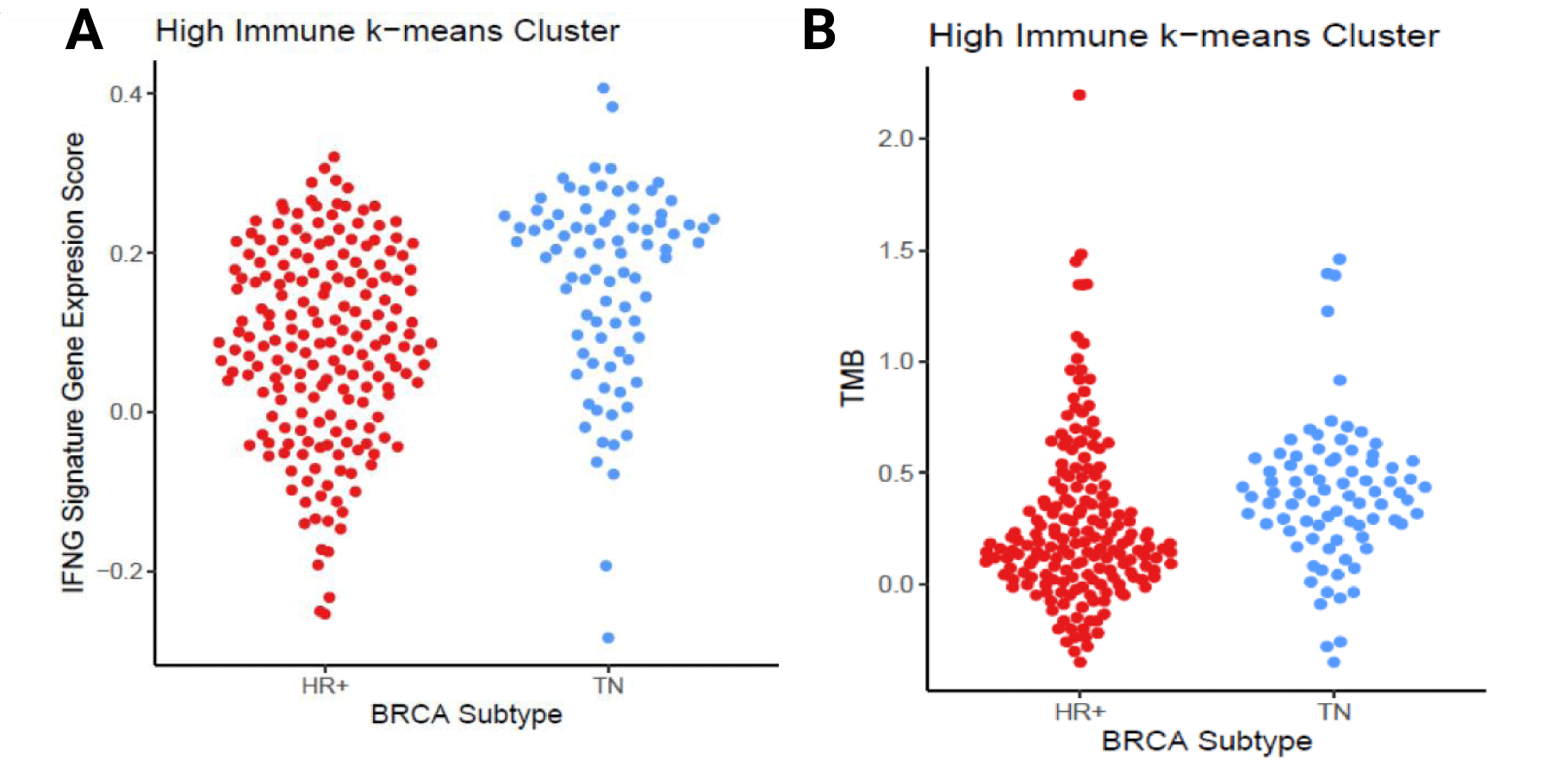
ICI efficacy is also associated with CD8+ T cells tumor infiltration and IFN-γ gene expression patterns in melanoma [35] and lung [6]. We used a gene signature that contains four genes (i.e., IFNγ, CD274, LAG3, and CXCL9; IFNγ+ signature), previously used to stratify urothelial and non-small-cell lung carcinoma cancer patient response to anti-PD-L1 durvalumab treatment to evaluate the immune-high and immune-low tumors [6]. When evaluating HR+HER2- and TN breast cancer, the mean of the IFNγ+ signature score was higher in TN than HR+HER2- (0.08 vs. -0.047; Welch’s 2 sided T-test p = 5.9x10-13), but the ranges of the scores were overlapping (TN: [-0.32, 0.41]; HR+: [-0.37, 0.32]). Similarly, when evaluating the immune-high group of HR+HER2- and TN tumors, the means of the IFNγ+signature scores are different, though less statistically significant (Welch’s 2 sided T-test, p-value = 2.2x10-6; figure 4A). Notably, when testing the IFNγ+ signature score between immune high and low cluster tumors, immune high tumors had significantly higher signatures scores irrespective of the breast cancer subtype (HR+HER2- Mann-Whitney test, p-value = 6.3x10-62, TN Mann-Whitney test, p-value = 1.3x10-18).
Tumor mutations generate neo-antigens and immune infiltrate can increase with an adaptive immune response to these neo-antigens increasing immune infiltrate. Tumor mutational burden also can determine response to ICI that is tumor-type agnostic [36]. To determine whether tumor mutational load differed between breast cancer subtypes, we calculated each tumor’s Tumor Mutational Burden (TMB). When evaluating the immune high group, the TN tumors contained a significantly higher TMB than the HR+HER2- tumors (Mann-Whitney test, p-value = 3.2X10-15; figure 4B). The difference in the median mutational burden between immune-high and immune-low was also statistically significant (Mann-Whitney test p-value = 1.2X10-5) with the immune high tumors having more TMB than the immune low tumors. The relationship between somatic mutations and the IFNγ+ signature shows a moderate correlation (Spearman’s ρ = 0.33), indicating that the immune response in breast cancer may in part be driven by somatic mutation derived neoantigens. Furthermore, evaluation of other mutation types (insertion, missense, nonsense) similarly showed significant differences between immune-high HR+HER2- and TN and immune-low HR+HER2- and TN (all p < 0.001, data not shown).
Fibroblast population associated with worse prognosis in breast cancer are higher in HR+HER2- tumors in both the immune-high and immune-low clusters.
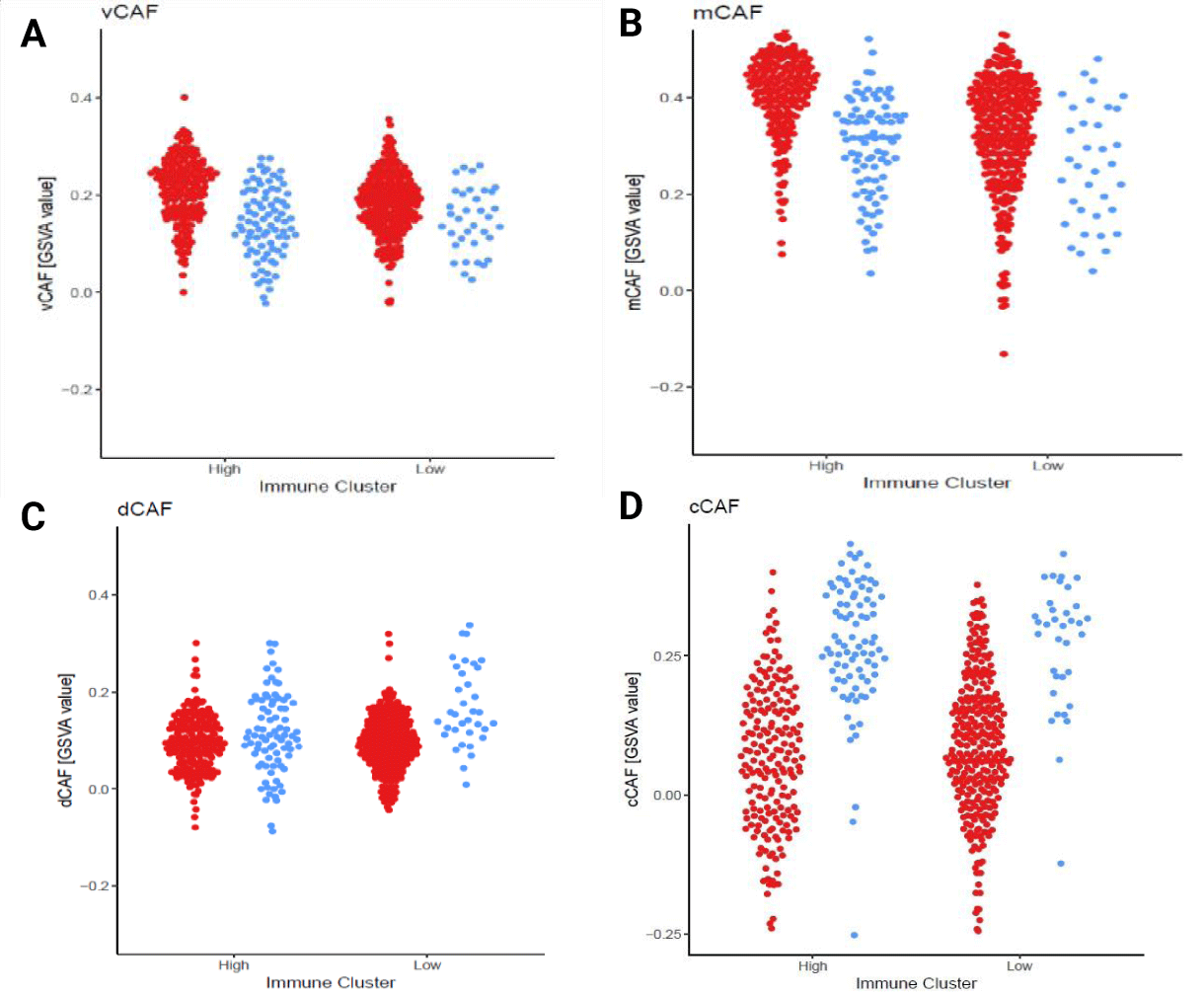
Many different populations of cancer-associated fibroblasts have been identified in breast cancer. Four populations that have been associated with prognosis in breast cancer in the TCGA include cell cycle related (cCAF), developmental (dCAF), extracellular matrix (mCAF), and vascular (vCAF) cancer associated fibroblasts. The vCAF signature was an independent prognostic indicator of developing metastatic disease. In the immune-high populations, the HR+HER2- tumors expressed higher vCAF fibroblast signature than the immune high TN tumors (p-value = 1.7x10-11). In the immune-low, the same trend was seen but there a less significant difference (p-value=7.8x10-3) (Figure 5A). The immune-high HR+HER2- tumors also had higher mCAF expression than the TN tumors (p-value=2.1x10-13). This mCAF signature has also been associated with increased risk of disseminated disease (Figure 5B) [28]. TN tumors had higher cCAF and dCAF expression than HR+HER2- tumors in both immune-high and immune-low populations (Figure 5(C,D)). These fibroblast types are not associated with tumor outcome [28]. There was an association between higher vCAF and lower TH1 tumor immune infiltrate in both the HR+HER2- immune-high (ρ = -0.6 p-value < 2.2x10-16) and HR+HER2- immune-low (ρ = -0.41 p-value = 6.5x10-13) and the TN immune-high (ρ = -0.36 p-value = 0.00065) but not in TN immune-low (ρ = -0.18 p-value = 0.3) (Supplementary figure 6(A,B)). There was also a significant negative association of fibroblasts to TH1 immune infiltrate in both the HR+HER2- immune-high (ρ = -0.63 p-value < 2.2x10-16) and HR+HER2- immune-low (ρ = -0.45 p-value = 1.6x10-15) as well as TN immune-high (ρ = -0.42 p-value = 6.9x10-5) and TN immune-low (ρ = -0.44 p-value = 0.0075) (Supplementary figure 6(C,D)).
Discussion
Expression analysis has identified subtypes of human breast cancer and can be used to guide treatment [37] but the differences in immune environment between the subtypes and how to address them in breast cancer remains unknown with relatively few breast cancer patients benefitting from immune based therapies. Evaluating immune gene clustering in TN and HR+HER2- breast cancer, the breast cancers did not stratify into canonical subtypes but rather divided into immune-high and immune-low tumors containing both subtypes. These immune-high and immune-low groups could be seen in two independent datasets, the TCGA and METABRIC [38,39]. Evaluation of the immune-high TN and HR+HER2- tumors demonstrated that they did not have significant differences in several common signatures associated with response to ICI and that the HR+HER2- immune high tumors contained both luminal A and luminal B tumors and the best and worst prognostic immune environments. The largest differences in expression between immune high TN and HR+HER2- tumors were that fibroblasts, specifically mCAF and vCAF that are associated with worse prognosis in breast cancer and were highest in HR+HER2- tumors while expression of Th1 and Th2 genes were highest in TN tumors.
We compared several signatures that are associated with increased immune response and improved response to ICIs including CD8+ and Treg signatures and the IFNγ signature [40]. The IFNγ+ gene signature can identify tumors that respond to anti-PD-L1 ICI durvalumab in 304 NSCLC patients and 103 urothelial cell cancers independent of PD-L1 expression [6]. The CD8+ and Treg signatures have been associated with improved response to immune checkpoint inhibitors in non-small cell lung cancer and urothelial cancer [6,7]. However, while all of these signatures were higher in the immune-high breast tumors, there were no significant differences between HR+HER2- and TN immune-high tumors. In immune-low tumors both CD8+, and Treg signatures were higher in TN compared to HR+HER2- (Figure 3D). There was no difference between CD8+ and Treg signatures between immune-high HR+HER2- and TN tumors and we found HR+HER2- and TN immune-high tumors to have similar ranges of IFNγ+ signature GSVA scores (Figure 4A). In immunogenic tumors such as melanoma or NSCLC, neo-antigens were associated with T cell mediated anti-tumor responses. Increased mutational load predicted a higher rate of response to anti-CTLA4 in melanoma and anti-PD-1 in lung cancer [41,42]. There was higher TMB in TN verses HR+HER2- tumors (p-value = 3.16X10-15). Furthermore, comparing the immune high TN and HR+HER2- tumors, there was more TMB in the TN immune high tumors (p-value = 3x10-5). This might explain the increased TH1 and TH2 immune infiltrate in the TN immune high tumors as compared to the HR+HER2- immune high tumors (Figure 4B) explaining the increased adaptive immune reactivity in TN immune-high tumors.
Six common immune environments can be seen across 33 diverse tumor types and determine tumor prognosis and response to therapy [27], with the C3 immune environment having the best clinical outcome. In this study, there was only six immune high TN tumors with a C3 immune environment therefore the majority of immune-high C3 tumors were HR+HER2-. Most immune-high TN tumors were the C2 immune subtype (Figure 2Bi). The C6 tumors predict the worst prognosis with increased TGF-β expression and are associated with increased tumor cell migration and activation of immune suppression from the tumor stroma in breast cancer [43,44]. Only HR+HER2- immune-high tumors had the C6 immune environment (Figure 2Bii). In the immune-high group, HR+HER2- breast cancer contained the best prognosis (C3) and worse prognosis (C6) tumors supporting that there are differences between immune-high TN and HER2+HER2- tumors that may explain why they have differential prognosis and response to ICI. We observed two major differences between HR+HER2- and TN within the immune-high tumors, TN immune-high tumors were enriched for Th1 and Th2 T cells while HR+HER2- immune-high tumors were enriched for fibroblasts, and TN immune-high tumors had increased TMB as compared to immune-high HR+HER2- tumors (Figure 3C,4B).
The increased expression of CAF in HR+HER2- immune-high breast tumors may suggest a mechanism that prevents these tumors from developing an appropriate anti-tumor immune response. The interaction between the tumor and stroma is important in determining the breast cancer subtype that develops [45,46]. The tumor stroma further can determine metastatic potential [47], resistance to chemotherapy [48], and modulating the immune response to the tumor [46]. When looking at breast cancer as a whole, expression of stromal and fibroblast components have been associated with worse prognosis including PDGFRA, PDGFRB, CXCL1, CXCL14, CD10, and CD36 [49]. CAFs have been associated with mesenchymal stem cells and inducing a wound healing response associated with immunosuppression [50]. In breast cancer, CAF expression has been correlated with decreased CD8+ T cells and increased macrophages [51]. Of interest, in a recent study of human breast cancer samples, four CAF subtypes were identified and the vascular (vCAF) and mesenchymal (mCAF) subtypes associated with developing metastatic disease [28]. In the TCGA dataset, both of those subtypes of CAF are significantly increased in the HR+HER2- breast tumors as compared to the TN breast tumors and there is higher significance in the immune-high population (Figure 5(A,B)). In this study we were unable to evaluate whether CAFs induce immunosuppression in immune-high HR+HER2- tumors but further evaluation of CAF in patients that do not respond to ICI is needed.
There are a few limitations in this study. The dataset lacks adequate tumor stage information thus we could not investigate the relationship between our findings and the stage of breast cancer.
Conclusion
Previous studies have shown that more advanced tumors have decreased immune infiltrate and increased immunosuppression [52]. Most importantly, with the lack of equivalent -omics and outcome data on ICI treated tumors, we could not evaluate the efficacy of ICIs on immune-high and -low tumors or compare ICI outcomes between HR+HER2- and TN tumors. While there have been some data showing benefit in HR+HER2- patients receiving neoadjuvant pembrolizumab with chemotherapy, the magnitude of response was 24.3% pCR compared to 64.8% pCR with addition of pembrolizumab to chemotherapy in TN patients [19,20]. Still, the biomarkers that predict response to ICI evaluated showed no significant differences between HR+HER2- and TN in the immune-high group: HR+HER2- and TN have similar expression of CD8+ and Treg expression while immune high TN does have higher TMB. We do find immune-high HR+HER2- tumors have increased CAF while immune-high TN tumors have increased helper T cells and increased expression of T cell modification genes. Further studies will need to be performed to determine whether these differences may affect how HR+HER2- and TN immune-high tumors respond to ICI.
Acknowledgement
We would like to thank Vanessa Hassle for her work on this project. Brian Fox, Josh Beilke, and Angela Davies for their comments on this manuscript. SES was supported by NIH T32 CA00951528, ITHS NIH KL2TR000421L2, and the Providence Foundation. MLD was supported by the Helen B. Slonaker Endowed Professor for Cancer Research, ACS Clinical Research Professor and a Komen Scholar Award.
Ethics approval and consent to participate
Not applicable
Consent for publication
Not applicable
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Authors’ contributions
FS analyzed and interpreted the data sets. WBC and JD aided in interpreting and discussing the data. SS interpreted the data and wrote the manuscript. MLD and KN contributing interpreting the data and revising the manuscript. All authors read and approved the final manuscript.
Competing interests
Authors received research funding, are employed by, and/or own stocks for the following organizations: Celgene (WC, JDA, KN, FS, MLD), Epithany (MLD), EMD Sorono (MLD), Precigen (MLD and SS), Barvarian Nordisk (MLD), Veanna (MLD and SS), IMV Inc (SS), Stanford, Burnham, Prebys (SS), and Pfizer (MLD).
References
- Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, Bryant J, Wolmark N. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004 Dec 30;351(27):2817-26. doi: 10.1056/NEJMoa041588. Epub 2004 Dec 10. PMID: 15591335.
- Johansson ALV, Trewin CB, Hjerkind KV, Ellingjord-Dale M, Johannesen TB, Ursin G. Breast cancer-specific survival by clinical subtype after 7 years follow-up of young and elderly women in a nationwide cohort. Int J Cancer. 2019 Mar 15;144(6):1251-1261. doi: 10.1002/ijc.31950. Epub 2018 Dec 3. PMID: 30367449.
- Liedtke C, Rody A, Gluz O, Baumann K, Beyer D, Kohls EB, Lausen K, Hanker L, Holtrich U, Becker S, Karn T. The prognostic impact of age in different molecular subtypes of breast cancer. Breast Cancer Res Treat. 2015 Aug;152(3):667-73. doi: 10.1007/s10549-015-3491-3. Epub 2015 Jul 21. PMID: 26195120.
- Sheridan W, Scott T, Caroline S, Yvonne Z, Vanessa B, David V, et al. Breast cancer in young women: have the prognostic implications of breast cancer subtypes changed over time? Breast Cancer Res Treat. 2014;147(3): 617-29.
- Stanton SE, Disis ML. Clinical significance of tumor-infiltrating lymphocytes in breast cancer. J Immunother Cancer. 2016 Oct 18;4:59. doi: 10.1186/s40425-016-0165-6. PMID: 27777769; PMCID: PMC5067916.
- Higgs BW, Morehouse CA, Streicher K, Brohawn PZ, Pilataxi F, Gupta A, Ranade K. Interferon Gamma Messenger RNA Signature in Tumor Biopsies Predicts Outcomes in Patients with Non-Small Cell Lung Carcinoma or Urothelial Cancer Treated with Durvalumab. Clin Cancer Res. 2018 Aug 15;24(16):3857-3866. doi: 10.1158/1078-0432.CCR-17-3451. Epub 2018 May 1. PMID: 29716923.
- Conway JR, Kofman E, Mo SS, Elmarakeby H, Van Allen E. Genomics of response to immune checkpoint therapies for cancer: implications for precision medicine. Genome Med. 2018 Nov 29;10(1):93. doi: 10.1186/s13073-018-0605-7. PMID: 30497521; PMCID: PMC6264032.
- Pagès F, Kirilovsky A, Mlecnik B, Asslaber M, Tosolini M, Bindea G, Lagorce C, Wind P, Marliot F, Bruneval P, Zatloukal K, Trajanoski Z, Berger A, Fridman WH, Galon J. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J Clin Oncol. 2009 Dec 10;27(35):5944-51. doi: 10.1200/JCO.2008.19.6147. Epub 2009 Oct 26. PMID: 19858404.
- Hwang WT, Adams SF, Tahirovic E, Hagemann IS, Coukos G. Prognostic significance of tumor-infiltrating T cells in ovarian cancer: a meta-analysis. Gynecol Oncol. 2012 Feb;124(2):192-8. doi: 10.1016/j.ygyno.2011.09.039. Epub 2011 Oct 29. PMID: 22040834; PMCID: PMC3298445.
- Dieu-Nosjean MC, Antoine M, Danel C, Heudes D, Wislez M, Poulot V, Rabbe N, Laurans L, Tartour E, de Chaisemartin L, Lebecque S, Fridman WH, Cadranel J. Long-term survival for patients with non-small-cell lung cancer with intratumoral lymphoid structures. J Clin Oncol. 2008 Sep 20;26(27):4410-7. doi: 10.1200/JCO.2007.15.0284. PMID: 18802153.
- Denkert C, Loibl S, Noske A, Roller M, Müller BM, Komor M, Budczies J, Darb-Esfahani S, Kronenwett R, Hanusch C, von Törne C, Weichert W, Engels K, Solbach C, Schrader I, Dietel M, von Minckwitz G. Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J Clin Oncol. 2010 Jan 1;28(1):105-13. doi: 10.1200/JCO.2009.23.7370. Epub 2009 Nov 16. Erratum in: J Clin Oncol. 2010 Feb 1;28(4):708. PMID: 19917869.
- Stanton SE, Adams S, Disis ML. Variation in the Incidence and Magnitude of Tumor-Infiltrating Lymphocytes in Breast Cancer Subtypes: A Systematic Review. JAMA Oncol. 2016 Oct 1;2(10):1354-1360. doi: 10.1001/jamaoncol.2016.1061. PMID: 27355489.
- Mahmoud SM, Paish EC, Powe DG, Macmillan RD, Grainge MJ, Lee AH, Ellis IO, Green AR. Tumor-infiltrating CD8+ lymphocytes predict clinical outcome in breast cancer. J Clin Oncol. 2011 May 20;29(15):1949-55. doi: 10.1200/JCO.2010.30.5037. Epub 2011 Apr 11. PMID: 21483002.
- Loi S, Sirtaine N, Piette F, Salgado R, Viale G, Van Eenoo F, Rouas G, Francis P, Crown JP, Hitre E, de Azambuja E, Quinaux E, Di Leo A, Michiels S, Piccart MJ, Sotiriou C. Prognostic and predictive value of tumor-infiltrating lymphocytes in a phase III randomized adjuvant breast cancer trial in node-positive breast cancer comparing the addition of docetaxel to doxorubicin with doxorubicin-based chemotherapy: BIG 02-98. J Clin Oncol. 2013 Mar 1;31(7):860-7. doi: 10.1200/JCO.2011.41.0902. Epub 2013 Jan 22. PMID: 23341518.
- Adams S, Gray RJ, Demaria S, Goldstein L, Perez EA, Shulman LN, Martino S, Wang M, Jones VE, Saphner TJ, Wolff AC, Wood WC, Davidson NE, Sledge GW, Sparano JA, Badve SS. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol. 2014 Sep 20;32(27):2959-66. doi: 10.1200/JCO.2013.55.0491. PMID: 25071121; PMCID: PMC4162494.
- Nanda R, Chow LQ, Dees EC, Berger R, Gupta S, Geva R, Pusztai L, Pathiraja K, Aktan G, Cheng JD, Karantza V, Buisseret L. Pembrolizumab in Patients With Advanced Triple-Negative Breast Cancer: Phase Ib KEYNOTE-012 Study. J Clin Oncol. 2016 Jul 20;34(21):2460-7. doi: 10.1200/JCO.2015.64.8931. Epub 2016 May 2. PMID: 27138582; PMCID: PMC6816000.
- Pernas S, Tolaney SM, Winer EP, Goel S. CDK4/6 inhibition in breast cancer: current practice and future directions. Ther Adv Med Oncol. 2018 Jul 17;10:1758835918786451. doi: 10.1177/1758835918786451. PMID: 30038670; PMCID: PMC6050811.
- Rugo HS, Delord JP, Im SA, Ott PA, Piha-Paul SA, Bedard PL, Sachdev J, Le Tourneau C, van Brummelen EMJ, Varga A, Salgado R, Loi S, Saraf S, Pietrangelo D, Karantza V, Tan AR. Safety and Antitumor Activity of Pembrolizumab in Patients with Estrogen Receptor-Positive/Human Epidermal Growth Factor Receptor 2-Negative Advanced Breast Cancer. Clin Cancer Res. 2018 Jun 15;24(12):2804-2811. doi: 10.1158/1078-0432.CCR-17-3452. Epub 2018 Mar 20. PMID: 29559561.
- Cardo so F, et al LBA 21 Keyno te 756: Phase III study of neoadjuvant pembrolizumab or placebo and chemotherapy followed by adjuvant pembro or pbo + endocrine therapy for early stage high risk ER+/HER2 - breast cancer. Annals of Oncology. 2023;34:s126 0-s1.
- Schmid P, Cortes J, Pusztai L, McArthur H, Kümmel S, Bergh J, Denkert C, Park YH, Hui R, Harbeck N, Takahashi M, Foukakis T, Fasching PA, Cardoso F, Untch M, Jia L, Karantza V, Zhao J, Aktan G, Dent R, O'Shaughnessy J; KEYNOTE-522 Investigators. Pembrolizumab for Early Triple-Negative Breast Cancer. N Engl J Med. 2020 Feb 27;382(9):810-821. doi: 10.1056/NEJMoa1910549. PMID: 32101663.
- Sharma P, Siefker-Radtke A, de Braud F, Basso U, Calvo E, Bono P, Morse MA, Ascierto PA, Lopez-Martin J, Brossart P, Rohrberg K, Mellado B, Fischer BS, Meadows-Shropshire S, Abdel Saci, Callahan MK, Rosenberg J. Nivolumab Alone and With Ipilimumab in Previously Treated Metastatic Urothelial Carcinoma: CheckMate 032 Nivolumab 1 mg/kg Plus Ipilimumab 3 mg/kg Expansion Cohort Results. J Clin Oncol. 2019 Jul 1;37(19):1608-1616. doi: 10.1200/JCO.19.00538. Epub 2019 May 17. Erratum in: J Clin Oncol. 2019 Aug 10;37(23):2094. doi: 10.1200/JCO.19.01700. PMID: 31100038; PMCID: PMC6879315.
- Wang L, Saci A, Szabo PM, Chasalow SD, Castillo-Martin M, Domingo-Domenech J, Siefker-Radtke A, Sharma P, Sfakianos JP, Gong Y, Dominguez-Andres A, Oh WK, Mulholland D, Azrilevich A, Hu L, Cordon-Cardo C, Salmon H, Bhardwaj N, Zhu J, Galsky MD. EMT- and stroma-related gene expression and resistance to PD-1 blockade in urothelial cancer. Nat Commun. 2018 Aug 29;9(1):3503. doi: 10.1038/s41467-018-05992-x. PMID: 30158554; PMCID: PMC6115401.
- Sinn BV, Loibl S, Karn T, Untch M, Kunze CA, Weber KE, D Treue, K Wagner, CA Hanusch, F Klauschen, PA Fasching, J Huober, D-M Zahm, C Jackisch, J Thomalla, J-U Blohmer, M van Mackelenbergh, K Rhiem, B Felder, G von Minckwitz, N Burchardi, A Schneeweiss, C Denkert. Abstract PD5-05: Pre-therapeutic PD-L1 expression and dynamics of Ki-67 and gene expression during neoadjuvant immune-checkpoint blockade and chemotherapy to predict response within the GeparNuevo trial. Cancer Research. 2019;79(4_Supplement): PD5-05. doi: 10.1158/1538-7445.SABCS18-PD5-05.
- Zhu Z, Turner NC, Loi S, Andre F, Martin M, Dieras V, Karen A. Gelmon, Nadia Harbeck, Cathy Zhang, Joan Q. Cao, Zhengming Yan, Dongrui R. Lu, Ping Wei, Todd L. VanArsdale, Paul A. Rejto, Xin Huang, Hope S. Rugo, Sibylle Loibl, Massimo Cristofanilli, Richard S. Finn, Yuan Liu. Comparative biomarker analysis of PALOMA-2/3 trials for palbociclib. NPJ Precis Oncol. 2022;6(1):56. doi: 10.1038/s41698-022-00297-1.
- Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012 Oct 4;490(7418):61-70. doi: 10.1038/nature11412. Epub 2012 Sep 23. PMID: 23000897; PMCID: PMC3465532.
- Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018 Nov;28(11):1747-1756. doi: 10.1101/gr.239244.118. Epub 2018 Oct 19. PMID: 30341162; PMCID: PMC6211645.
- Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, Sivakumar IKA, Gentles AJ, Malhotra R, Farshidfar F, Colaprico A, Parker JS, Mose LE, Vo NS, Liu J, Liu Y, Rader J, Dhankani V, Reynolds SM, Bowlby R, Califano A, Cherniack AD, Anastassiou D, Bedognetti D, Mokrab Y, Newman AM, Rao A, Chen K, Krasnitz A, Hu H, Malta TM, Noushmehr H, Pedamallu CS, Bullman S, Ojesina AI, Lamb A, Zhou W, Shen H, Choueiri TK, Weinstein JN, Guinney J, Saltz J, Holt RA, Rabkin CS; Cancer Genome Atlas Research Network; Lazar AJ, Serody JS, Demicco EG, Disis ML, Vincent BG, Shmulevich I. The Immune Landscape of Cancer. Immunity. 2018 Apr 17;48(4):812-830.e14. doi: 10.1016/j.immuni.2018.03.023. Epub 2018 Apr 5. Erratum in: Immunity. 2019 Aug 20;51(2):411-412. doi: 10.1016/j.immuni.2019.08.004. PMID: 29628290; PMCID: PMC5982584.
- Bartoschek M, Oskolkov N, Bocci M, Lövrot J, Larsson C, Sommarin M, Madsen CD, Lindgren D, Pekar G, Karlsson G, Ringnér M, Bergh J, Björklund Å, Pietras K. Spatially and functionally distinct subclasses of breast cancer-associated fibroblasts revealed by single cell RNA sequencing. Nat Commun. 2018 Dec 4;9(1):5150. doi: 10.1038/s41467-018-07582-3. PMID: 30514914; PMCID: PMC6279758.
- Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013 Jan 16;14:7. doi: 10.1186/1471-2105-14-7. PMID: 23323831; PMCID: PMC3618321.
- Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017 Nov 15;18(1):220. doi: 10.1186/s13059-017-1349-1. PMID: 29141660; PMCID: PMC5688663.
- Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015 Apr 20;43(7):e47. doi: 10.1093/nar/gkv007. Epub 2015 Jan 20. PMID: 25605792; PMCID: PMC4402510.
- Gulbahce HE, Sweeney C, Surowiecka M, Knapp D, Varghese L, Blair CK. Significance of GATA-3 expression in outcomes of patients with breast cancer who received systemic chemotherapy and/or hormonal therapy and clinicopathologic features of GATA-3-positive tumors. Hum Pathol. 2013 Nov;44(11):2427-31. doi: 10.1016/j.humpath.2013.05.022. Epub 2013 Aug 30. PMID: 23998430.
- Barrueto L, Caminero F, Cash L, Makris C, Lamichhane P, Deshmukh RR. Resistance to Checkpoint Inhibition in Cancer Immunotherapy. Transl Oncol. 2020 Mar;13(3):100738. doi: 10.1016/j.tranon.2019.12.010. Epub 2020 Feb 27. PMID: 32114384; PMCID: PMC7047187.
- Morrison C, Pabla S, Conroy JM, Nesline MK, Glenn ST, Dressman D, Papanicolau-Sengos A, Burgher B, Andreas J, Giamo V, Qin M, Wang Y, Lenzo FL, Omilian A, Bshara W, Zibelman M, Ghatalia P, Dragnev K, Shirai K, Madden KG, Tafe LJ, Shah N, Kasuganti D, de la Cruz-Merino L, Araujo I, Saenger Y, Bogardus M, Villalona-Calero M, Diaz Z, Day R, Eisenberg M, Anderson SM, Puzanov I, Galluzzi L, Gardner M, Ernstoff MS. Predicting response to checkpoint inhibitors in melanoma beyond PD-L1 and mutational burden. J Immunother Cancer. 2018 May 9;6(1):32. doi: 10.1186/s40425-018-0344-8. PMID: 29743104; PMCID: PMC5944039.
- Karachaliou N, Gonzalez-Cao M, Crespo G, Drozdowskyj A, Aldeguer E, Gimenez-Capitan A, Teixido C, Molina-Vila MA, Viteri S, De Los Llanos Gil M, Algarra SM, Perez-Ruiz E, Marquez-Rodas I, Rodriguez-Abreu D, Blanco R, Puertolas T, Royo MA, Rosell R. Interferon gamma, an important marker of response to immune checkpoint blockade in non-small cell lung cancer and melanoma patients. Ther Adv Med Oncol. 2018 Jan 18;10:1758834017749748. doi: 10.1177/1758834017749748. PMID: 29383037; PMCID: PMC5784541.
- Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, Lu S, Kemberling H, Wilt C, Luber BS, Wong F, Azad NS, Rucki AA, Laheru D, Donehower R, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, Greten TF, Duffy AG, Ciombor KK, Eyring AD, Lam BH, Joe A, Kang SP, Holdhoff M, Danilova L, Cope L, Meyer C, Zhou S, Goldberg RM, Armstrong DK, Bever KM, Fader AN, Taube J, Housseau F, Spetzler D, Xiao N, Pardoll DM, Papadopoulos N, Kinzler KW, Eshleman JR, Vogelstein B, Anders RA, Diaz LA Jr. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017 Jul 28;357(6349):409-413. doi: 10.1126/science.aan6733. Epub 2017 Jun 8. PMID: 28596308; PMCID: PMC5576142.
- Dowsett M, Sestak I, Lopez-Knowles E, Sidhu K, Dunbier AK, Cowens JW, Ferree S, Storhoff J, Schaper C, Cuzick J. Comparison of PAM50 risk of recurrence score with oncotype DX and IHC4 for predicting risk of distant recurrence after endocrine therapy. J Clin Oncol. 2013 Aug 1;31(22):2783-90. doi: 10.1200/JCO.2012.46.1558. Epub 2013 Jul 1. PMID: 23816962.
- Cline MS, Craft B, Swatloski T, Goldman M, Ma S, Haussler D, Zhu J. Exploring TCGA Pan-Cancer data at the UCSC Cancer Genomics Browser. Sci Rep. 2013 Oct 2;3:2652. doi: 10.1038/srep02652. PMID: 24084870; PMCID: PMC3788369.
- Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Doug Speed, Andy G. Lynch, Shamith Samarajiwa, Yinyin Yuan, Stefan Gräf, Gavin Ha, Gholamreza Haffari, Ali Bashashati, Roslin Russell, Steven McKinney, METABRIC Group, Anita Langerød, Andrew Green, Elena Provenzano, Gordon Wishart, Sarah Pinder, Peter Watson, Florian Markowetz, Leigh Murphy, Ian Ellis, Arnie Purushotham, Anne-Lise Børresen-Dale, James D. Brenton, Simon Tavaré, Carlos Caldas, Samuel Aparicio. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486(7403):346-52.
- Havel JJ, Chowell D, Chan TA. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer. 2019 Mar;19(3):133-150. doi: 10.1038/s41568-019-0116-x. PMID: 30755690; PMCID: PMC6705396.
- Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS, Hollmann TJ, Bruggeman C, Kannan K, Li Y, Elipenahli C, Liu C, Harbison CT, Wang L, Ribas A, Wolchok JD, Chan TA. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014 Dec 4;371(23):2189-2199. doi: 10.1056/NEJMoa1406498. Epub 2014 Nov 19. Erratum in: N Engl J Med. 2018 Nov 29;379(22):2185. doi: 10.1056/NEJMx180040. PMID: 25409260; PMCID: PMC4315319..
- Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, Zappasodi R, Maeda Y, Sander C, Garon EB, Merghoub T, Wolchok JD, Schumacher TN, Chan TA. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015 Apr 3;348(6230):124-8. doi: 10.1126/science.aaa1348. Epub 2015 Mar 12. PMID: 25765070; PMCID: PMC4993154.
- Wiercinska E, Naber HP, Pardali E, van der Pluijm G, van Dam H, ten Dijke P. The TGF-β/Smad pathway induces breast cancer cell invasion through the up-regulation of matrix metalloproteinase 2 and 9 in a spheroid invasion model system. Breast Cancer Res Treat. 2011 Aug;128(3):657-66. doi: 10.1007/s10549-010-1147-x. Epub 2010 Sep 5. PMID: 20821046.
- de Kruijf EM, Dekker TJA, Hawinkels LJAC, Putter H, Smit VTHBM, Kroep JR, Kuppen PJK, van de Velde CJH, Ten Dijke P, Tollenaar RAEM, Mesker WE. The prognostic role of TGF-β signaling pathway in breast cancer patients. Ann Oncol. 2013 Feb;24(2):384-390. doi: 10.1093/annonc/mds333. Epub 2012 Sep 28. PMID: 23022998.
- Sidaway P. Breast cancer: Fibroblast subtypes alter the microenvironment. Nat Rev Clin Oncol. 2018 May;15(5):264-265. doi: 10.1038/nrclinonc.2018.37. Epub 2018 Mar 6. PMID: 29508858.
- Wallace JA, Li F, Leone G, Ostrowski MC. Pten in the breast tumor microenvironment: modeling tumor-stroma coevolution. Cancer Res. 2011 Feb 15;71(4):1203-7. doi: 10.1158/0008-5472.CAN-10-3263. Epub 2011 Feb 8. PMID: 21303970; PMCID: PMC3075554.
- Finak G, Bertos N, Pepin F, Sadekova S, Souleimanova M, Zhao H, Chen H, Omeroglu G, Meterissian S, Omeroglu A, Hallett M, Park M. Stromal gene expression predicts clinical outcome in breast cancer. Nat Med. 2008 May;14(5):518-27. doi: 10.1038/nm1764. Epub 2008 Apr 27. PMID: 18438415.
- Farmer P, Bonnefoi H, Anderle P, Cameron D, Wirapati P, Becette V, André S, Piccart M, Campone M, Brain E, Macgrogan G, Petit T, Jassem J, Bibeau F, Blot E, Bogaerts J, Aguet M, Bergh J, Iggo R, Delorenzi M. A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat Med. 2009 Jan;15(1):68-74. doi: 10.1038/nm.1908. Epub 2009 Jan 4. Erratum in: Nat Med. 2009 Feb;15(2):220. Wirapati, Pratyakasha [corrected to Wirapati, Pratyaksha]. PMID: 19122658.
- Brechbuhl HM, Finlay-Schultz J, Yamamoto TM, Gillen AE, Cittelly DM, Tan AC, Sams SB, Pillai MM, Elias AD, Robinson WA, Sartorius CA, Kabos P. Fibroblast Subtypes Regulate Responsiveness of Luminal Breast Cancer to Estrogen. Clin Cancer Res. 2017 Apr 1;23(7):1710-1721. doi: 10.1158/1078-0432.CCR-15-2851. Epub 2016 Oct 4. PMID: 27702820; PMCID: PMC5378660.
- Kalluri R. The biology and function of fibroblasts in cancer. Nat Rev Cancer. 2016 Aug 23;16(9):582-98. doi: 10.1038/nrc.2016.73. PMID: 27550820.
- Costa A, Kieffer Y, Scholer-Dahirel A, Pelon F, Bourachot B, Cardon M, Sirven P, Magagna I, Fuhrmann L, Bernard C, Bonneau C, Kondratova M, Kuperstein I, Zinovyev A, Givel AM, Parrini MC, Soumelis V, Vincent-Salomon A, Mechta-Grigoriou F. Fibroblast Heterogeneity and Immunosuppressive Environment in Human Breast Cancer. Cancer Cell. 2018 Mar 12;33(3):463-479.e10. doi: 10.1016/j.ccell.2018.01.011. Epub 2018 Feb 15. PMID: 29455927.
- Harano K, Wang Y, Lim B, Seitz RS, Morris SW, Bailey DB, Hout DR, Skelton RL, Ring BZ, Masuda H, Rao AUK, Laere SV, Bertucci F, Woodward WA, Reuben JM, Krishnamurthy S, Ueno NT. Rates of immune cell infiltration in patients with triple-negative breast cancer by molecular subtype. PLoS One. 2018 Oct 12;13(10):e0204513. doi: 10.1371/journal.pone.0204513. PMID: 30312311; PMCID: PMC6193579.
Content Alerts
SignUp to our
Content alerts.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.