Biology Group. 2024 September 20;5(9):1140-1151. doi: 10.37871/jbres2002.
Metabolic Engineering of the purine metabolic pathway in brewer's yeast to reduce the Purine components in a Beer System
Jun Liu1-4 and Jian Lu1,3,4*
2Sericultural and Agri-Food Research Institute, Guangdong Academy of Agricultural Sciences, Key Laboratory of Functional Foods, Ministry of Agriculture and Rural Affairs, Guangdong Key Laboratory of Agricultural Products Processing, Guangzhou 510610, China
3National Engineering Research Center of Cereal Fermentation and Food Biomanufacturing, Jiangnan University, Wuxi, 214122, China
4Jiangsu Provincial Research Center for Bioactive Product Processing Technology, Jiangnan University, Wuxi, 214122, China
- Metabolic engineering
- Brewer's yeast
- Purine metabolic pathway
Abstract
Beer is one of the most popular beverages in the world. However, beer contains excessive amounts of purines, which can increase the risk of hyperuricaemia and gout. During beer fermentation, a key reason is that the purines are not completely absorbed by yeast, which is due to the insufficient ability of purine salvage pathway in beer yeast. In order to improve the absorption capacity of yeast for purines, this study overexpressed four key genes in the purine metabolism pathway (HPT1, APT1, FCY2, and PRM15) in the commercial beer yeast strain SC4, and obtained three yeast engineering strains, namely M1 (SC4 HPT1:: APT1), M2 (M1:: FCY2), and M3 (M2:: PRM15). The expression of purine metabolism-related genes was examined at the transcriptional level, and the expression levels of HPT1, APT1, FCY2 and PRM15 genes were increased by 3.50-fold, 2.37-fold, 2.07-fold and 4.37-fold, respectively, and the overexpression of FCY2 and PRM15 increased the expression levels of HPT1 and APT1 by 3.50-fold and 2.37-fold, respectively. The expression of FCY2 and PRM15 made HPT1 and APT1 more active, indicating that the ability to absorb purines was further enhanced. Applying engineering yeast M3 to 12-degree wort fermentation, the results shown that the free purine base content is reduced by 53.35%, which provides a theoretical basis for reducing the purine content in beer.
Introduction
Beer is a low-alcohol beverage with barley malt as its main raw material, which is popular with consumers all over the world because of its rich nutrition, unique flavor and pleasant taste [1,2]. Numerous studies have shown that there is a high dependency between beer and gout, and that gout patients who drink beer may induce gout attacks [3]. During saccharification, most of the Ribonucleic Acid (RNA) and Deoxyribonucleic Acid (DNA) in barley malt are degraded to produce soluble nitrogenous substances such as purine nucleotides, purine nucleosides and purine bases under the action of ribonucleic acid enzyme system and thermodynamics, of which free purine bases, such as adenine and guanine, account for about 6%, which is an important driving force for yeast proliferation. During beer fermentation, purine substances are mainly assimilated and absorbed by brewer's yeast. In the early stage of fermentation, yeast assimilates free purine bases through the purine remediation synthesis pathway to synthesize intracellular nucleic acids and achieve rapid proliferation in order to shorten the growth retardation period and fermentation time. When there is a lack of free purine bases in the wort, the yeast will not be able to start the purine remediation synthesis pathway, and can only synthesize the required purines through the de novo synthesis pathway using sugars and amino acids, which consumes a large amount of energy, and thus greatly affects its proliferation capacity and fermentation performance. Nucleosides cannot be transported to the body through the yeast cell membrane [4], beer yeast cannot utilize purine nucleosides, and purine nucleosides from raw material sources, such as barley malt, end up residing in the body of the brew, resulting in excessive purine analogues in the beer.
Yeast genetically engineered stain and genetically modified crops have not gained consumer acceptance in the food and alcoholic beverage industry. Self-cloning and non-selfcloning techniques are two methods of genetic improvement based on the origin of genes [5]. The use of yeast ‘gene self-cloning’ to modify the yeast genome [6] has eliminated the fear of consumers of products labelled with genetically engineered stains, as there is no introduction of heterologous DNA and the Cartagena Protocol on Biosafety states that it will not be controlled [2]. In the field of yellow wine industrial yeast, Wu D, et al. [7] used self-cloning technology to modify the arginine metabolic pathway of industrial strain N85, and successfully reduced the EC content of harmful substances in yellow wine by 55.3%. In beer industrial yeasts, which are more widely used, researchers used seif-cloning to increase glutathione content, reduce proteinase A for better foam stability, enhance yeast flocculation and improve acetaldehyde reduction [8-11]. Therefore, gene self-cloning technique has important applications in constructing industrial yeast strains with excellent characteristics. There are no reports on the use of self-cloning technology to modify low purine brewer's yeast.
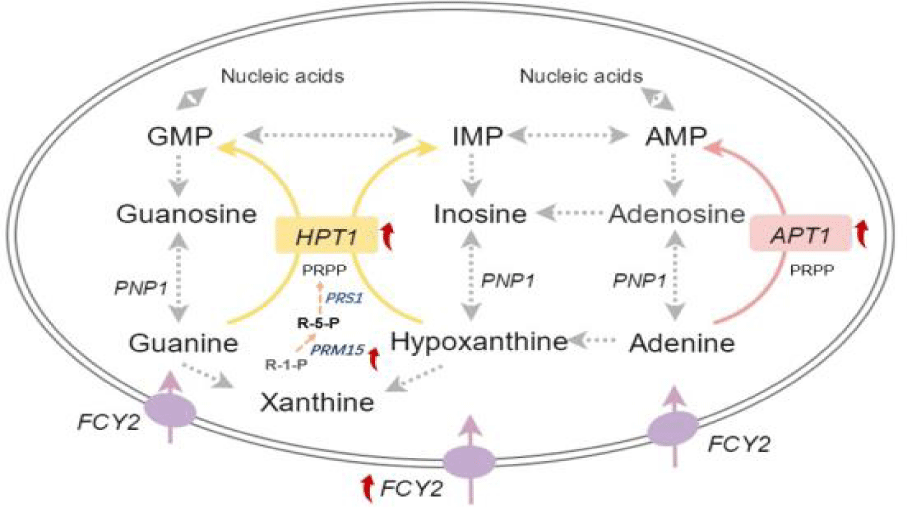
In the yeast purine salvage pathway, the two most critical enzymes are Hypoxanthine-guanine Phosphoribosyl Transferase (HPT) [12] and Adenine Phosphoribosyl Transferase (ART) [13], which catalyse the reaction of free purine bases with PRPP to produce the corresponding purine nucleotides to shorten the growth retardation period. The APT genes consist of APT1 and APT2, and APT2 is an analogue produced by the whole genome duplication of APT1, and the cell growth is normal after null mutation normal, and when both APT1 and APT2 are null mutated, cells show growth defects [13]. Phosphoribomutase (PRM15) [14] catalyses the interconversion of ribulose-1-phosphate and ribulose-5-phosphate and has some phosphoglucose translocase activity, but the main activity in vivo is that of phosphoribomutase, which contributes to ribose recycling in the pentose phosphate pathway, and can relieve IMP from the feedback inhibitory effect on PRS. Purine-cytosine permease (FCY2) is a proton transporter that takes up adenine, guanine, hypoxanthine and cytosine from extracellular sources as raw materials for the remedial synthesis pathway [15]. Although FCY2 is functionally similar to FCY21 and FCY22, FCY21 or FCY22 cannot complement the null mutant function of FCY2 [15,16]. Therefore, in order to improve the ability of brewer's yeast to assimilate and take up free purines, and to reduce the purine content of beer while increasing yeast proliferation and improving fermentation performance, HPT1, APT1, PRM15, and FCY2 could all serve as potential targets for metabolic engineering modification.
In our previous study, the addition of PNPs at the saccharification stage significantly increased the content of free purine bases in wort for yeast proliferation during fermentation, but the yeast's ability to assimilate free bases was found to be limited, and a certain amount of free purine bases were still present in the finished beer. In order to further reduce the purine content in beer, this study used "selfcloning technology" to strengthen the purine uptake pathway of brewer's yeast, and constructed highly efficient expression assemblies of HPT1, APT1, FCY2, and PRM15 genes with a strong yeast promoter, and then gradually transformed them into brewer's yeast starter strain SC4, so that the yeast engineered bacteria were obtained with a low purine content. The fermentation performance and purine uptake capacity of the engineered yeast were investigated.
Materials and Methods
Yeast strain, plasmids, and media
All plasmids and strains used in this study are listed in table 1. The parent strain, beer yeast SC4, is an industrial Ale-brewing strain and Escherichia coli JM109 strains, conserved in our laboratory, was used. Vector pMD19-T (simple) was purchased from Takara (Dalian, China) and plasmids pUG6 and YP26-tef-gpd were obtained from our laboratory.
| Table 1: Microbial strains and plasmids used in this study. | ||
| Strains and plasmids | Relevant characteristics | Source |
| SC4 | commercial yeast strain | Our lab |
| PUG6 | containing -loxP-KanMX-loxP- expression cassette | Our lab |
| pY26 | containing TEF1p-ADH1t and GDP1p- CYC1t | Our lab |
| pSH65 | containing Zeocin ble and Cre recombinase-expressed genes | Our lab |
| pYPGE15 | containing PGK1 promoter | Our lab |
| JM109 | E.coli strain | Our lab |
| M1 | SC4::TEF1p-HPT1-ADH1 and GPD1p-APT1-CYC1t | This study |
| M2 | M1:: PGK1p-FCY2 | This study |
| M3 | M2:: PGK1p-PRM15 | This study |
E. coli JM109 was grown at 37 °C in Luria-Bertani (LB) medium (composed of 10 g/L NaCl, 10 g/L tryptone, and 5 g/L yeast extract) supplemented with ampicillin (100 μg/mL) to select positive E. coli transformants. Yeast Extract Peptone Dextrose (YEPD) medium (composed of 10 g/L yeast extract, 20 g/L peptone, and 20 g/L glucose) was used to cultivate the yeast strains. YEPD plates were supplemented with 100 μg/mL filter-sterilized G418 antibiotic (Sangon, Shanghai, China) to select for yeast transformants harboring the KanMX gene. All the solid media used in this study was prepared with 20 g/L agar. Polymerase Chain Reaction (PCR) primers were synthesized by Sangon Biotech (Shanghai, China) and are listed in table 2. The genomic DNA of S. cerevisiae was extracted using a Ezup Column Yeast Genomic DNA Purification Kit (Sangon, Shanghai, China). The plasmids had G418 selection markers derived from plasmid pUG6.
| Table 2: Primers used in this study. | |
| Name | 5′ → 3′ DNA sequence |
| HPT1f | agaattgttaattaaagatctTCATTGCTTGTGTTCCTGCTCA |
| HPT1r | aactccggaccgcggagatctATGTCGGCAAACGATAAGCAA |
| APT1f | caggaattcgatatcaagcttATGTCTATAGCAAGTTATGCCCAAGA |
| APT1r | gtcgacggtatcgataagcttTCATTTTTTCAACGCTTCCTTTT |
| URA3-L1 | GCATTAGGTCCCAAAATTTGT |
| URA3-L2 | gcacacggtgtggtgggcccCAGGAAACAGCTATGACCATGATT |
| URA3-R1 | cctttgttacttcttccgccCGAGGTCGACGGTATCGATAA |
| URA3-R2 | CGAGGTCGACGGTATCGATA |
| KKANA1f | cctttgttacttcttccgccATTTAGGTGACACTATAGAACGCGG |
| KANA1r | cctttgttacttcttccgccTAATACGACTCACTATAGGGAGACCG |
| ble1 | ATCCGTCGAGTGGGTGGT |
| ble2 | AATGACCGACCAAGCGAC |
| CYC1f | gcacacggtgtggtgggcccCAGGAAACAGCTATGACCAT |
| ADH1r | ttctatagtgtcacctaaatCGAGGTCGACGGTATCGATAA |
| FCY2-L1 | GCCTCAAGGAACTAAATATCAGT |
| FCY2-L2 | tattataacgatttataagaATTTAGGTGACACTATAGAACGCGG |
| FCY2-r1 | gatcctgtcattgttataatATGTTGGAAGAGGGAAATAATG |
| FCY2-r2 | GAAAAGAACGCAACAAATATCAACCCC |
| KANA2f | tattataacgatttataagaATTTAGGTGACACTATAGAACGCGG |
| KANA2r | gaagtcaggaatctaaaataTAATACGACTCACTATAGGGAGACCG |
| PGK1p1f | ccctatagtgagtcgtattaTATTTTAGATTCCTGACTTCAACTCAAGA |
| PGK1p1r | ttatttccctcttccaacatATTATAACAATGACAGGATCCCTGTTT |
| PRM15-L1 | CCCATCTCCGACGAGGAATT |
| PRM15-L2 | ggagaaggggatgacaacgaATTTAGGTGACACTATAGAACGCGG |
| PRM15-r1 | atcctgtcattgttataatATGTTGCAAGGAATT |
| PRM15-r2 | TGCAGCAGCAGTAGCTCTAGCGA |
| KANA3f | ggagaaggggatgacaacgaATTTAGGTGACACTATAGAACGCGG |
| KANA3r | gaagtcaggaatctaaaataTAATACGACTCACTATAGGGAGACCG |
| PGK1p2f | ccctatagtgagtcgtattaTATTTTAGATTCCTGACTTCAACTCAAGA |
| PGK1p2r | tctaaaattccttgcaacatATTATAACAATGACAGGATCCCTGTTT |
RNA preparation and cDNA synthesis
Yeast cells were harvested by centrifugation (4 °C, 5400 rpm, 1 min) after beer fermentation for 48h and immediately stored in liquid nitrogen until RNA extraction. The cells were ground in liquid nitrogen, and total RNA was prepared using TRIzol reagent (Invitrogen, Carlsbad, CA). The RNA was quantified and checked in a Nanodrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, DE) at 260 nm and 280 nm. The integrity of the isolated RNA was verified with an automated electrophoresis system (Bio-Rad, Hercules, CA). cDNA was synthesized from 5 g of total RNA using the PrimeScript RT reagent kit (TaKaRa, Dalian, China).
Construction of recombinant vectors to express HPT1 and APT1
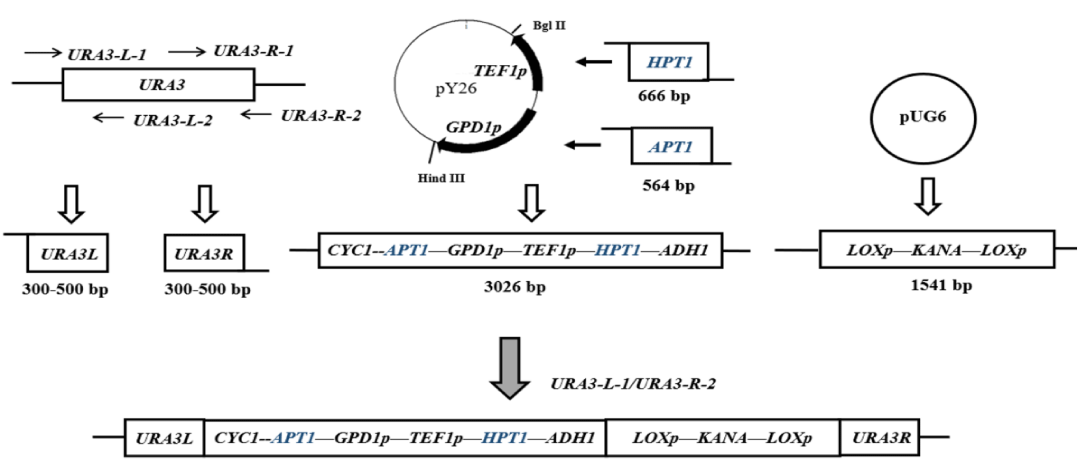
The HPT1 and APT1 efficient expression assembly was constructed as shown in figure 1. Using the genome of brewer's yeast SC4 as a template, the HPT1 and APT1 genes with homology arms were obtained by PCR amplification using primers HPT1f/HPT1r and APT1f/APT1r with Prime STAR DNA polymerase. Then, using restriction endonuclease Bgl II and Hind III, respectively, pY26 was enzymatically ligated to obtain the pY26-HPT1-APT1 recombinant plasmid; the upstream and downstream homology arms were amplified using the primers URA3-L1/ URA3-L2 and URA3-R1/ URA3-R2, respectively, to obtain URA3-L and URA3-R; and using plasmid PUG6 as a template. Using plasmid PUG6 as the template, the KanMX resistance marker fragment with loxp was amplified using primers KANA1f/ KANA1r; finally, the fragments were fused according to the ClonExpress MultiS One Step Multiple Fragment Ligation Kit from Goldvector to obtain the high-efficiency expression assemblies of HPT1 and APT1, which were then subjected to PCR and sequencing for identification. The successfully validated HPT1 and APT1 high-efficiency expression components were integrated into the yeast genome by yeast lithium acetate transformation, and positive transformants were screened on YPD plates containing 80 mg/L G418 antibiotic. After the positive transformants were grown, the recombinant engineered bacteria were identified by PCR and sequenced using URA3-L1/ URA3-R2 primers. Finally, the pSH65 plasmid with Cre recombinase was transferred into the yeast engineering strain, and the G418 resistance was excised by cyclisation using the Cre/loxP system, which facilitated the transformation of the subsequent expression components.
Construction of recombinant vectors to express FCY2 and PRM15
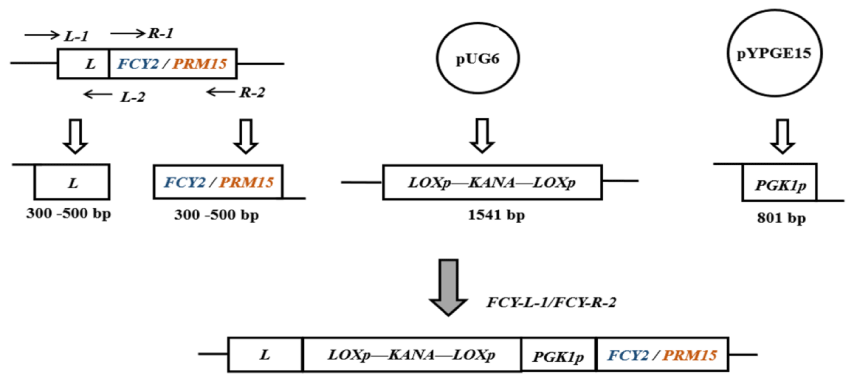
FCY2 or PRM13 high efficiency expression assemblies are shown in figure 2. The strong promoter PGK1p was inserted in front of the modified genes and loxp-KanMX-loxp was added as a screening marker. The genome of brewer's yeast SC4 was used as a template for PCR amplification of the upstream and downstream homology arm fragments L and R (part of the FCY2 gene) using Prime STAR DNA polymerase and primers FCY2-L1/ FCY2-L2 and FCY2-R1/ FCY2-R2; plasmid PUG6 was used as a template for PCR amplification of the upstream and downstream homology arm fragments L and R (part of the FCY2 gene) using primers KANA2f/ KANA2r to amplify the KanMX resistance marker fragment with loxP; plasmid pYPGE15 as a template, and primers PGK1p1f / PGK1p1r to amplify the PGK1p promoter fragment; and finally, fusion of fragments according to the ClonExpress MultiS One Step Multi-Fragment Ligation Kit from Goldvein, to obtain the FCY2 The FCY2 high-efficiency expression components were fused according to the ClonExpress MultiS One Step Multiple Fragment Ligation Kit, and the components were identified by PCR and sequencing. The successfully validated FCY2 high-efficiency expression component was integrated into the yeast genome by yeast lithium acetate transformation, and the positive transformants were screened on YPD plates containing 80 mg/L G418 antibiotic. After the positive transformants were grown, the recombinant engineered bacteria were identified by PCR and sequenced using primers FCY2-L1 / FCY2-R2. Finally, the pSH65 plasmid with Cre recombinase was transferred into the yeast engineering strain, and the G418 resistance was excised by cyclisation using Cre/loxP system for the next genetic transformation operation. The genome of brewer's yeast SC4 was used as a template for PCR amplification of PRM15-L1/ PRM15-L2 and PRM15-R1/ PRM15-R2 using primers to obtain the upstream and downstream homology arm fragments L and R. The plasmid PUG6 was used as a template for amplification of the KanMX resistance marker fragments with loxP using the primers KANA3f/ KANA3r; and the plasmid pYPGE15 was used as the template, and the PGK1p promoter fragment was amplified using primers PGK1p2f / PGK1p2r; finally, the fragments were fused according to the ClonExpress MultiS One Step Multi-fragment Ligation Kit from KingVision to obtain the PRM15 high-efficiency expression assembly, which was then subjected to PCR and sequencing for identification. The PRM15 high-efficiency expression assembly, which was successfully validated, was integrated into the yeast genome by yeast lithium acetate transformation, and positive transformants were screened on YPD plates containing 80 mg/L G418 antibiotic. After the positive transformants were grown, the recombinant engineered bacteria were identified by PCR and sequenced using primers PRM15-L1 / PRM15-R2. Finally, the pSH65 plasmid with Cre recombinase was transferred into the yeast engineering strain and the G418 resistance was excised by cyclisation using Cre/loxP system.
qRT-PCR: All primer sequences were designed for ACT1, HPT1, APT1, FCY2 and PRM15 by using Beacon Designer 7.0 (Table 3). The efficiencies and specificities of the primers were tested in dilution experiments and by constructing melting curves, respectively. The real-time quantitative PCR (qRT-PCR) experiments were performed with the SYBR premix ExTaq kit (TaKaRa) with the synthesized cDNA as the template. The cycling parameters for the PCR were as follows: preincubation at 95 °C for 30 s, 40 cycles of amplification at 95 °C for 5 s and 60 °C for 20 s, and a final cooling step at 50 °C for 30 s. The reactions were conducted in a Stepone real-time PCR instrument (Thermo Fisher Scientific, USA). All experiments were performed in biological replicates with an independent measurement of each sample, and mean values were used for further calculations. The fold changes were determined via the cycle threshold (2ΔCt) method and normalized to the results for the ACT1 gene. Measurement of purines, and the main components in the model beer system.
| Table 3: Oligonucleotide primers used for qRT-PCR. | ||
| Gene | Primer name | 5′ → 3′ DNA sequence |
| ACT1 | ACT1-F | ATGGATTCTGAGGTTGCTGC |
| ACT1-R | ACCGACGATAGATGGGAAGAC | |
| HPT1 | HPT1-F | CTGAGCAGGAACACAAG CAATGA |
| HPT1-R | GACACCCGTACCACACTTCA | |
| APT1 | APT1-F | GGAATCCCGTGGGTTCTTGT |
| APT1-R | AACGTTGGAACCTGCTGGAA | |
| FCY2 | FCY2-F | GTAAGACAACAGTTTCCAGACAATT |
| FCY2-R | GGAACTAAAGGGGTATGAACAA | |
| PRM15 | PRM15-F | ATGGTGAATGAAGGCTCCG |
| PRM15-R | TGATCTCGATAGTTGGGCAAT | |
Beer fermentation: The mashing procedure was as follow: 10 min at 53 °C, 60 min at 63 °C, 10 min at 72 °C, and 10 min at 78 °C. The heating rate for temperature ramp-up was 1 °C per min; moreover, 100 g of barley malt was milled and mashed with a water-to-malt ratio of 4:1 in a mashing bath (BGT-8A, Bioer). The resulting mash was then filtered to separate the wort, and the residual mash was washed with warm water. Subsequently, the wort, to which 0.3 g/L hops (Magnum, α-acid 12-14%, Germany) had been added was allowed to boil for 1 h. After the wort cooled to room temperature, it was filtered through a filter cloth, and the sugar content of the filtrate was adjusted to 12 Plato.
Yeast cells were precultured in 3 mL of 12 °Plato wort medium for 24 h at 28 °C with shaking at 220 rpm. These cells were then transferred to 30 mL of wort medium in a 150 mL conical flask and incubated for 12 h at 28 °C with shaking at 220 rpm. The yeast cells were then resuspended in wort to achieve a cell concentration of 2-3 × 108 cells/mL. Subsequently, 10 mL of this inoculum was transferred to a 500 mL conical flask containing 200 mL of wort medium, and incubated at 10ºC until completion, when the daily weight loss was below 0.2 g. The daily weight loss was calculated by measuring the weight of fermented liquid with the fermentation equipment. All fermentations were carried out in triplicates.
Determination of fermentation performance
Fermentation-performance indicators (wort density and ethanol content) were measured using a pycnometer (Chinese National Standard for Beer Analysis GB/T 4928-2008) and residual sugar was analyzed via the Fehling titration method. The real degree of fermentation (RDF) was calculated using equation.
where A denotes ethanol content (%), and Z represents wort density (%).
Determination of purine content
Purine compounds were analyzed using High Performance Liquid Chromatography (HPLC) (Agilent 1260, USA) [17].
Determination of amino acid content
The amino acid content of beer was determined using o-phthalaldehyde derivatisation coupled with HPLC [18].
Statistical Analyses
Raw data were further analyzed by Student’s test (*p < 0.05, **p < 0.01). Each experiment was performed in at least triplicate biological samples. Statistical analyses were performed using the statistical software SPSS (SPSS Inc., Chicago, IL, USA).
Results and Discussion
Construction of recombinant brewer's yeast engineering strains
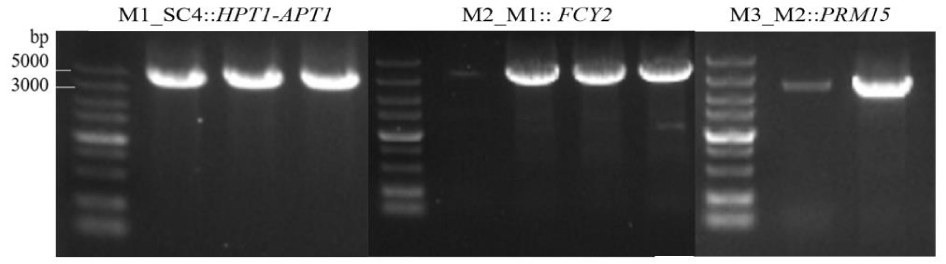
In our previous study, it was found that the addition of purine nucleoside phosphorylase into the saccharification process produces a large amount of free purines, and a significant portion of free purine bases remained in the beer at the end of the fermentation (p < 0.05), indicating that the yeast's ability to utilise free purines is limited. Therefore, in order to improve the ability of brewer's yeast to assimilate and absorb free purines, we chose to use HPT1, APT1, PRM15 and FCY2 as targets for metabolic engineering modification in this study. Firstly, using KanMX as a screening marker, we constructed a high-efficiency expression assembly of HPT1 and APT1 genes with a length size of 5033 bp, and after successful sequencing validation, we integrated them into the yeast SC4 genome by yeast lithium acetate transformation to obtain the yeast engineering strain M1 (SC4::HPT1-APT1). Using Cre/loxP knockout system, KanMX resistance was excised cyclically in M1, and then the constructed FCY2 high-efficiency expression assembly, with a length and size of 3,365 bp, was integrated into Saccharomyces cerevisiae M1 to obtain yeast engineering strain M2 (M1::FCY2). After excision of KanMX resistance, it was transferred into PRM15 high-efficiency expression component with length size 3031 bp, and the yeast engineered strain was named M3 (M2::PRM15), and finally the KanaMX resistance was recovered to eliminate the food safety risk caused by exogenous genes. As shown in figure 3 the size of electrophoresis bands was as expected, and the sequencing results were consistent with the electrophoresis results, indicating the successful construction of brewer's yeast engineering strain.
Effect of metabolic engineering modification on gene expression in brewer's yeast
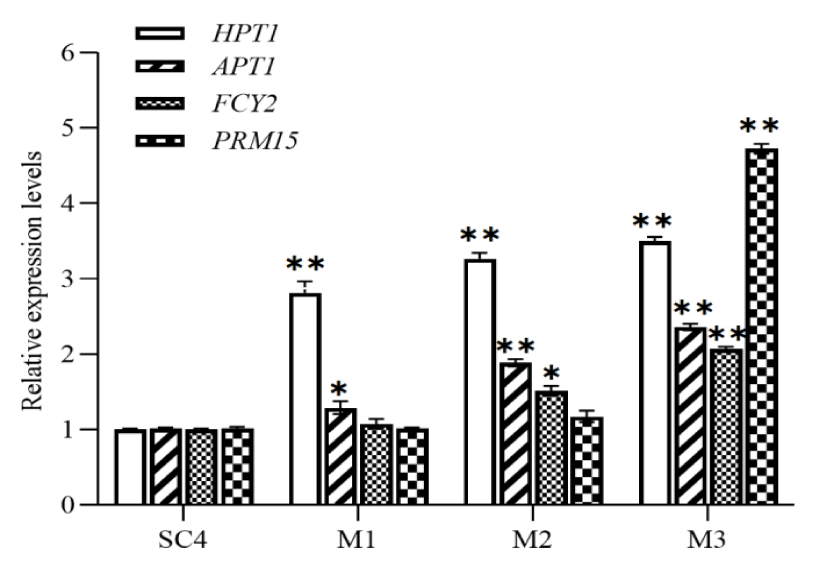
When wort is rich in free purines, yeast rapidly absorbs purine bases from the environment through the remedial synthesis pathway for its own proliferative metabolism in order to shorten the latency period. In the yeast purine salvage pathway, the two most critical enzymes are HPT [12] and APT [13], of which the APT gene consists of APT1 and APT2, and APT2 is the analogue produced by the whole-genome duplication of APT1, which grows the cell normally after the null mutation, when the APT1 and APT2 are both null mutated, cells show growth defects, indicating an APT1 key functional gene [13]. Highly efficient expression of HPT1 and APT1 in brewer's yeast SC4 using strong promoters TEF1p and GPD1p resulted in 2.87 ± 0.11-fold and 1.32 ± 0.17-fold increase in gene expression, respectively (Figure 4). FCY2 is a proton transporter that takes up adenine, guanine, hypoxanthine and cytosine from extracellular sources as raw materials for the remedial synthesis pathway [15]. To further increase the flux of free purines into yeast cells, FCY2 was further expressed efficiently in this study, with a 1.51 ± 0.15-fold increase in gene expression, and the expression of HPT1 and APT1 was elevated to 3.27 ± 0.25 and 1.9 ± 0.28-fold with the increase in the amount of purine bases entering the yeast cells. Ribulose 5-phosphate is an important raw material in the remedial synthesis pathway, and nucleic acid ribose translocase (PRM15) catalyses the interconversion between ribose 1-phosphate and ribose 5-phosphate, which contributes to the ribose recycling in the pentose phosphate pathway, and also disengages the feedback inhibitory effect of IMP on the PRPP synthase (PRS1), when overexpressed PRM15 expression was elevated to 4.37 ± 0.10-fold, and HPT1, APT1 and PRM15 increased 3.50 ± 0.14-fold, 2.37 ± 0.18-fold and 2.07 ± 0.09-fold, respectively. Among them, the bacterial difference of HPT1 gene in all engineered strains was significant (p < 0.01) compared with the control, and HPT1, APT1, FCY2 and PRM15 genes were significantly increased (p < 0.01) in engineered strain M3 compared with the control, when the overexpression of FCY2 and PRM15 caused HPT1 and APT1 to become more active, implying a further enhancement of the ability to absorb purines.
Effect of metabolic engineering modification on purine uptake by brewer's yeast
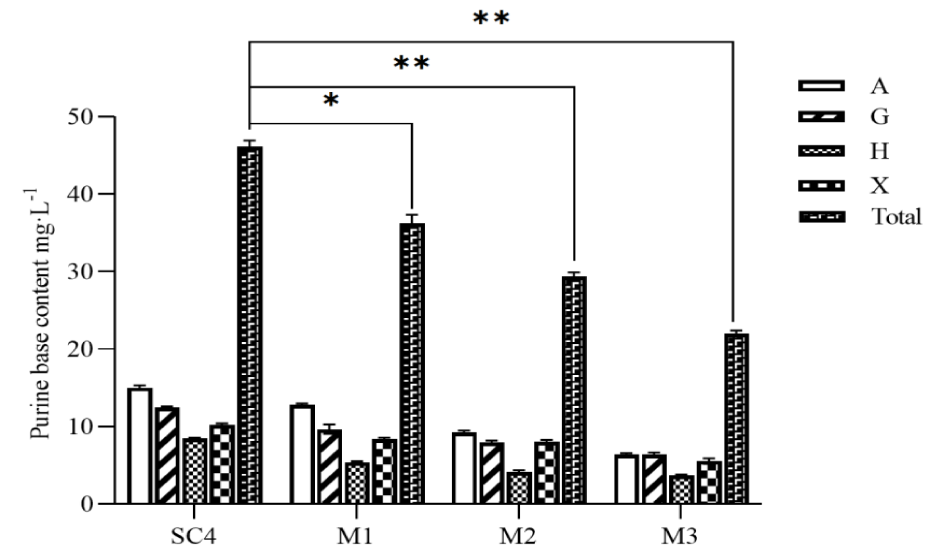
The free purine content of beer fermentation broth was shown in figure 5, and the purine content of the engineered bacteria M1, M2, and M3 were all significantly decreased (p < 0.05) compared with the control SC4 (47.07 ± 1.09 mg/L), in which the yeast engineered strain M3 was the strongest in uptake of purine, with the residual purine content of 21.96 ± 0.45 mg/L (p < 0.01), with a reduction ratio of 53.35%, followed by M1 and M2 with 36.25 ± 1.09 mg/L (p < 0.05) and 29.42 ± 0.47 mg/L (p < 0.01) respectively, with a reduction ratio of 22.99% and 37.50%. It was shown that increasing the expression of HPT1, APT1, FCY2 and PRM15 enhanced the uptake of free purine in yeast.
Effect of metabolic engineering modifications on the fermentation performance of brewer's yeast
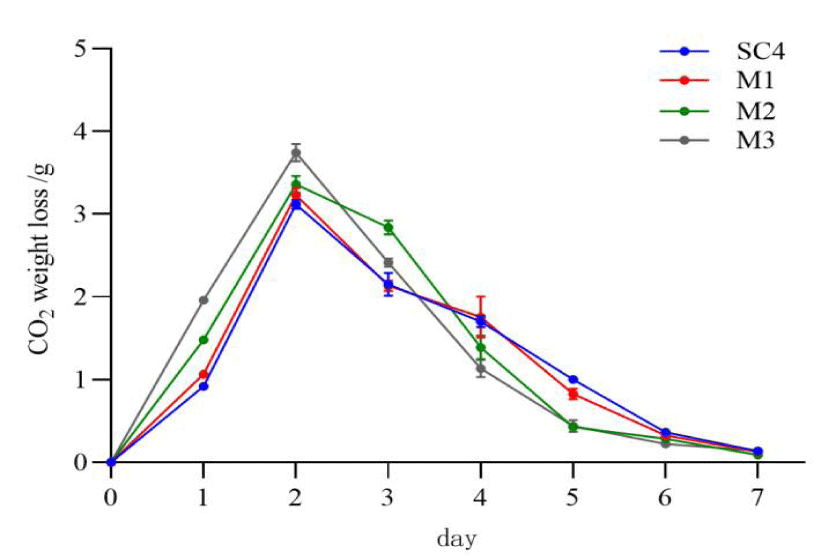
Yeast engineered bacteria fermentation curves: After inoculating in wort, the yeast makes use of the nutrients such as carbon and ammonia in wort to grow and multiply by aerobic respiration, yeast cells produce ethanol and CO2 by anaerobic fermentation [19]. The fermentation rate and activity of yeast can be directly judged by the weight loss of CO2. In this study, CO2 weight loss curves for fermentation of low purine brewer's yeast engineered bacteria were determined, as shown in figure 6. The fermentation times of engineered bacteria M1, M2 and M3 were the same as that of the control SC4, which was 7 d. In terms of the overall CO2 weight loss, the fermentation performance of M3 was better than that of M1 and M2, and the CO2 weight loss curves reached a maximum on the second day, which was an increase of 12.68% compared to the control (p < 0.05).
Physico-chemical characteristics of fermentation broth: Brewer's yeast fermentation degree is one of the most important fermentation characteristics, which reflects the ability of brewer's yeast to utilise sugar. The physico-chemical characteristics of the beer brewed with the engineered yeasts are shown in table 4. The fermentation broths of the engineered stain M1, M2, and M3 showed different degrees of increase in alcohol, fermentation degree, and total acid, and a decrease in pH, compared to the fermentation broths of the starting strain SC4. vol, an increase of 5.72% (p < 0.05); fermentation degree increased from 73.35% to 74.70%, an increase of 1.84% (p < 0.05); and the differences in pH and total acid were significant (p < 0.05).It indicated that the metabolic engineering modification of HPT1, APT1, FCY2 and PRM15 genes significantly improved the fermentation performance of brewer's yeast.
| Table 4: Basic physico-chemical characteristics of beers. | ||||
| Physico-chemical characteristics | SC4 | M1 | M2 | M3 |
| Alcohol %/ vol | 4.76 ± 0.07 | 4.77 ± 0.11 | 4.83 ± 0.09 | 5.03 ± 0.07* |
| Real fermentation degree/ % | 73.35 ± 0.28 | 73.65 ± 0.47 | 73.90 ± 0.31 | 74.70 ± 0.42* |
| Original Gravity / % w/w | 12.27 ± 0.12 | 12.25 ± 0.19 | 12.14 ± 0.23 | 12.33 ± 0.26 |
| pH | 4.30 ± 0.01 | 4.28 ± 0.01 | 4.27 ± 0.01 | 4.21 ± 0.01* |
| Total acid /mL/100 mL | 2.48 ± 0.09 | 2.49 ± 0.14 | 2.51 ± 0.16 | 2.56 ± 0.12* |
| Data are averages of three independent experiments, and error bars represent ± SD (*p < /em> < 0.05, **p < /em>< 0.01). | ||||
Free amino acid content of beer fermentation broth: The variation of free amino acid content in beer fermentation broth was determined as shown in table 5. The total amino acid content in the fermentation broths of the beer yeast engineered bacteria M1, M2 and M3 were 567.77, 560.70 and 551.46 mg/L, respectively, which were reduced by 4.03%, 5.22% and 6.78%, respectively, compared with the control SC4 (591.60 mg/L). Based on the preferential consumption of amino acids during beer fermentation, amino acids can be classified into four groups, A, B, C and D. At the end of fermentation, group A (Thr, Met, Lys, Ser, Leu and Arg) was assimilated at a faster rate than group B (Ile, Phe, His, Asp and Val), followed by group C (Tyr, Pro and Glu), and the least uptake of amino acids was in group D (Gly and Ala) [20,21], Lys, His, Arg and Leu were all derived from the corresponding amino acids in the wort, which were hardly synthesised from saccharides and had a significant effect on yeast metabolism and proliferation, Pro was the amino acid with the lowest assimilation by the yeast, and Ala, Gly and Pro were the least preferred amino acids by the yeast, which were basically the same in the fermentation broth. Similar results were obtained in the present study re, Pro was the highest in the fermentation broth of all strains, the metabolically modified yeast engineered yeast fermentation broth was essentially unchanged (p > 0.05) for Ala, Gly and Pro compared to the control, with a significant (p < 0.05) decrease in Lys, His, Arg and Leu. In Thomas and Ingledew [22] it was found that the addition of Glu to ethanol production shortened the fermentation time, indicating that Glu improves fermentation performance. Free amino acid uptake is closely related to the degree of fermentation and alcohol content [21]. The significant (p < 0.05) decrease in Glu in group C belonging to the present study may be due to the improved fermentation performance. In conclusion, metabolic engineering modification enhanced the uptake of amino acids by yeast during fermentation and obtained better fermentation performance.
| Table 5: Detection of amino acids in beer samples fermented by different strains. | ||||
| Amino acids/mg/L | SC4 | M1 | M2 | M3 |
| Asp < /td> | 8.76 ± 0.26 | 4.46 ± 0.04* | 5.85 ± 0.09* | 7.13 ± 0.49 |
| Glu | 23.75 ± 0.66 | 18.59 ± 0.63* | 16.67 ± 0.37* | 15.19 ± 0.95* |
| Ser | 0.58 ± 0.01 | 0.56 ± 0.01 | 0.10 ± 0.00** | 0.07 ± 0.00** |
| His | 16.75 ± 0.80 | 9.04 ± 0.75* | 7.85 ± 0.19* | 7.95 ± 0.60* |
| Gly | 23.95 ± 0.91 | 23.59±0.42 | 23.83 ± 0.57 | 23.18 ± 0.43 |
| Thr | 14.93 ± 0.82 | 14.32 ± 1.11 | 14.47 ± 0.25 | 14.17 ± 0.21 |
| Arg | 1.95 ± 0.03 | 1.41 ± 0.02* | 1.54 ± 0.07 | 1.66 ± 0.06* |
| Ala | 10.72 ± 0.43 | 10.20 ± 0.09 | 10.09 ± 0.11 | 10.61 ± 0.12 |
| Tyr | 23.61 ± 0.50 | 22.91 ± 0.61 | 21.03 ± 1.25 | 20.35 ± 1.08 |
| Cys-s | 1.88 ± 0.03 | 1.77 ± 0.10 | 1.71 ± 0.13 | 1.10 ± 0.00* |
| Val | 23.33 ± 0.87 | 23.73 ± 0.46 | 23.61 ± 1.73 | 23.17 ± 0.27 |
| Met | 2.56 ± 0.08 | 2.24 ± 0.03 | 2.34 ± 0.09 | 2.05 ± 0.05 |
| Phe | 3.74 ± 0.04 | 3.97 ± 0.17 | 3.19 ± 0.05 | 3.25 ± 0.04 |
| Ile | 1.81 ± 0.07 | 2.06 ± 0.04 | 3.96 ± 0.10 | 2.69 ± 0.07 |
| Leu | 2.62 ± 0.07 | 2.42 ± 0.05 | 1.51 ± 0.28* | 0.92 ± 0.06* |
| Lys | 1.30 ± 0.01 | 0.91 ± 0.01* | 0.34 ± 0.00* | 0.22 ± 0.01* |
| Pro | 429.45 ± 8.48 | 425.60 ± 6.25 | 422.61 ± 6.08 | 417.83 ± 9.01 |
| Tatol | 591.60 | 567.77 | 560.70 | 551.46 |
| Data are averages of three independent experiments, and error bars represent ± SD (*p < /em> < 0.05, **p < /em> < 0.01). | ||||
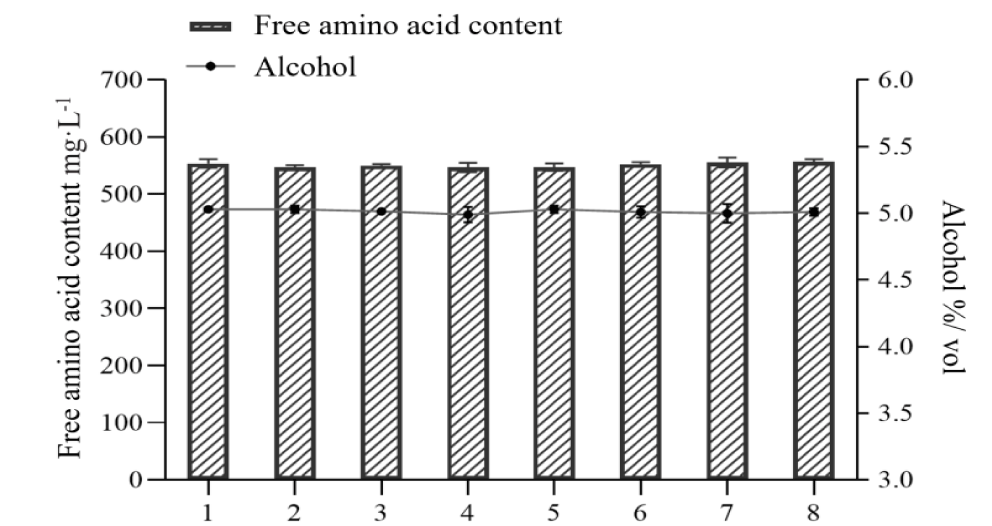
Effect of metabolic engineering modifications on genetic stability of brewer's yeast: Due to the existence of repair mechanisms in microorganisms themselves, strains with stable genetic characteristics can only be used in industrial production [23]. Therefore, the genetic stability of the brewer's yeast engineered strain M3 was further verified. Successive passages (8 generations) of M3 were cultured to determine the purine, alcohol and amino acid contents in the beer fermentation broth. As shown in table 6 and figure 7, the engineering strain M3 had good genetic stability, the total free amino acid content was maintained at 552.00 mg/L, the alcohol content was maintained at 5.00% vol, and the average reduction rate of free purine was 53.64%, indicating that the recombinant yeast strain had good genetic stability.
| Table 6: Effect of continuous transmission of yeast engineering on purine content in beer fermentation broth. | |||
| Generations | Free purine content /mg/L | Purine reduction rate /% | |
| SC4 | M3 | ||
| 1 | 47.07 ± 1.47 | 21.96 ± 0.45 | 53.35 |
| 2 | 47.28 ± 1.21 | 21.37 ± 0.53 | 54.80 |
| 3 | 46.54 ± 1.03 | 21.04 ± 0.24 | 54.79 |
| 4 | 45.65 ± 1.05 | 21.38 ± 0.37 | 53.17 |
| 5 | 46.34 ± 0.94 | 21.15 ± 0.42 | 54.36 |
| 6 | 48.32 ± 1.31 | 23.28 ± 0.35 | 51.82 |
| 7 | 47.54 ± 1.24 | 21.24 ± 0.43 | 55.32 |
| 8 | 48.15 ± 1.17 | 23.31 ± 0.22 | 51.59 |
| Averages | 47.11 | 21.84 | 53.64 |
| Data are averages of three independent experiments ± SD. | |||
Conclusion
In this study, we constructed efficient expression assemblies of HPT1, APT1, FCY2 and PRM15 genes using a strong yeast promoter, and gradually transformed them into a brewer's yeast starter strain to obtain a low-purine yeast engineering strain, and examined the ability of the engineering strain to reduce the purine content of beer. We obtained the brewer's yeast engineering strain M3 by transforming the starting strain SC4 through gene self-cloning. The results of laboratory triangular flask fermentation of Engineering Strain M3 showed that, compared with the starting strain SC4, the fermentation performance and purine absorption ability of Engineering Strain M3 increased by 5.72% in alcohol content, 1.84% in degree of fermentation, and the ability to utilize free purine increased by 53.35%; The results of laboratory triangular flask fermentation of Engineering Strain M3 indicated that the fermentation performance of engineering strain M3 and its capacity of utilizing free purine increased by 53.35%. The engineering strain M3 had good genetic stability, and no response mutation occurred in 8 generations.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors are aware and agree with the submission of this manuscript.
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Author contribution
Jun Liu: Conceptualization, Methodology, Data curation, Writing - original draft. Jian Lu: Reviewing and Editing, Supervision. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the GuangDong Basic and Applied Basic Research Foundation (2023A1515111173); the Jinying zhiguang Project of GDAS (R2023PY-JG016); the Program of Introducing Talents of Discipline to Universities (grant number 111-2-06) and the Collaborative Innovation Center of Jiangsu Modern Industrial Fermentation.
References
- Rodhouse L, Carbonero F. Overview of craft brewing specificities and potentially associated microbiota. Crit Rev Food Sci Nutr. 2019;59(3):462-473. doi: 10.1080/10408398.2017.1378616. Epub 2017 Oct 4. PMID: 28910550.
- Petruzzi L, Corbo M R, Sinigaglia M, Bevilacqua A. Brewer's yeast in controlled and uncontrolled fermentations, with a focus on novel, nonconventional, and superior strains. Food Reviews International. 2016;32(4):341-363. doi: 10.1080/87559129.2015.1075211.
- Ragab G, Elshahaly M, Bardin T. Gout: An old disease in new perspective - A review. J Adv Res. 2017 Sep;8(5):495-511. doi: 10.1016/j.jare.2017.04.008. Epub 2017 May 10. PMID: 28748116; PMCID: PMC5512152.
- Lee W J, Prentice N. Utilization of nucleosides and nucleobases by the lager yeast, Saccharomyces Carlsbergensis. Journal of the American Society of Brewing Chemists. 1987;45(4):128-131. doi: 10.1094/asbcj-45-0128.
- Akada R. Genetically modified industrial yeast ready for application. J Biosci Bioeng. 2002;94(6):536-44. doi: 10.1016/s1389-1723(02)80192-x. PMID: 16233347..
- Fischer S, Procopio S, Becker T. Self-cloning brewing yeast: a new dimension in beverage production. European Food Research and Technology. 2013;237(6):851-863. doi: 10.1007/s00217-013-2092-9.
- Wu D, Li X, Lu J, Chen J, Zhang L, Xie G. Constitutive expression of the DUR1,2 gene in an industrial yeast strain to minimize ethyl carbamate production during Chinese rice wine fermentation. FEMS Microbiol Lett. 2016 Jan;363(1):fnv214. doi: 10.1093/femsle/fnv214. Epub 2015 Nov 3. PMID: 26538578.
- Ogata T, Kobayashi M, Gibson BR. Pilot-scale brewing using self-cloning bottom-fermenting yeast with high SSU1 expression. Journal of the Institute of Brewing. 2013;119(1-2):17-22. doi: 10.1002/jib.61.
- Kusunoki K, Ogata T. Construction of self-cloning bottom-fermenting yeast with low vicinal diketone production by the homo-integration of ILV5. Yeast. 2012 Oct;29(10):435-42. doi: 10.1002/yea.2922. Epub 2012 Oct 5. PMID: 23038161.
- Wang Z Y, He X P, Liu N, Bo-Run Z. Construction of self-cloning industrial brewing yeast with high-glutathione and low-diacetyl production. International Journal of Food Science & Technology. 2008;43(6):989-994. doi: 10.1111/j.1365-2621.2007.01546.x.
- Wang ZY, Wang JJ, Liu XF, He XP, Zhang BR. Recombinant industrial brewing yeast strains with ADH2 interruption using self-cloning GSH1+CUP1 cassette. FEMS Yeast Res. 2009 Jun;9(4):574-81. doi: 10.1111/j.1567-1364.2009.00502.x. Epub 2009 Apr 1. PMID: 19341381.
- Woods RA, Roberts DG, Friedman T, Jolly D, Filpula D. Hypoxanthine: guanine phosphoribosyltransferase mutants in Saccharomyces cerevisiae. Mol Gen Genet. 1983;191(3):407-12. doi: 10.1007/BF00425755. PMID: 6355764.
- Alfonzo JD, Crother TR, Guetsova ML, Daignan-Fornier B, Taylor MW. APT1, but not APT2, codes for a functional adenine phosphoribosyltransferase in Saccharomyces cerevisiae. J Bacteriol. 1999 Jan;181(1):347-52. doi: 10.1128/JB.181.1.347-352.1999. PMID: 9864350; PMCID: PMC103569.
- Xu YF, Létisse F, Absalan F, Lu W, Kuznetsova E, Brown G, Caudy AA, Yakunin AF, Broach JR, Rabinowitz JD. Nucleotide degradation and ribose salvage in yeast. Mol Syst Biol. 2013 May 14;9:665. doi: 10.1038/msb.2013.21. PMID: 23670538; PMCID: PMC4039369.
- Wagner R, Straub ML, Souciet JL, Potier S, de Montigny J. New plasmid system to select for Saccharomyces cerevisiae purine-cytosine permease affinity mutants. J Bacteriol. 2001 Jul;183(14):4386-8. doi: 10.1128/JB.183.14.4386-4388.2001. PMID: 11418581; PMCID: PMC95330.
- Nelissen B, Mordant P, Jonniaux JL, De Wachter R, Goffeau A. Phylogenetic classification of the major superfamily of membrane transport facilitators, as deduced from yeast genome sequencing. FEBS Lett. 1995 Dec 18;377(2):232-6. doi: 10.1016/0014-5793(95)01380-6. PMID: 8543057.
- Li H, Liu F, Hao J. Determination of purines in beer by HPLC using a simple and rapid sample pretreatment. Journal of the American Society of Brewing Chemists. 2015;73(2):137-142. doi: 10.1094/ASBCJ-2015-0409-01.
- Cigić I, Vodošek T, Košmerl T, M. Strlič. Amino acid quantification in the presence of sugars using HPLC and pre-column derivatization with 3-MPA/OPA and FMOC-Cl. Acta Chimica Slovenica. 2008;55(3):660-664.
- Capece A, Romaniello R, Pietrafesa A, Siesto G, Pietrafesa R, Zambuto M, Romano P. Use of Saccharomyces cerevisiae var. boulardii in co-fermentations with S. cerevisiae for the production of craft beers with potential healthy value-added. Int J Food Microbiol. 2018 Nov 2;284:22-30. doi: 10.1016/j.ijfoodmicro.2018.06.028. Epub 2018 Jul 2. PMID: 29990636.
- Jones M, Pierce J S. Absorption of amino acids from wort by yeasts. Journal of the Institute of Brewing. 1964;70(4):307-315. doi: 10.1002/j.2050-0416.1964.tb01996.x.
- Lei H, Zheng L, Wang C, Zhao H, Zhao M. Effects of worts treated with proteases on the assimilation of free amino acids and fermentation performance of lager yeast. Int J Food Microbiol. 2013 Feb 1;161(2):76-83. doi: 10.1016/j.ijfoodmicro.2012.11.024. Epub 2012 Dec 6. PMID: 23279816.
- Thomas KC, Ingledew WM. Fuel alcohol production: effects of free amino nitrogen on fermentation of very-high-gravity wheat mashes. Appl Environ Microbiol. 1990 Jul;56(7):2046-50. doi: 10.1128/aem.56.7.2046-2050.1990. PMID: 2202254; PMCID: PMC184558.
- Verbelen PJ, Dekoninck TM, Van Mulders SE, Saerens SM, Delvaux F, Delvaux FR. Stability of high cell density brewery fermentations during serial repitching. Biotechnol Lett. 2009 Nov;31(11):1729-37. doi: 10.1007/s10529-009-0067-5. Epub 2009 Jun 30. PMID: 19565190.
Content Alerts
SignUp to our
Content alerts.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.