Biology Group. 2024 September 20;5(9):1152-1163. doi: 10.37871/jbres2003.
Occult Hepatitis C Infection in Kidney Transplant and Hemodialysis Patients Identified five to Eight Years after Sustained Virological Response
Eliane Silva1,2*, Sara Marques1, Tereza Almeida1, Fabiana Neves1, Maria Dias Lopes3, José Filipe Santos3, Jorge Leitão3,4, Adélia Simão3,4 and Armando Carvalho3,4
2School of Medicine and Biomedical Sciences of the University of Porto (ICBAS-UP), Porto, Portugal
3Unidade Local de Saúde de Coimbra (ULSC), Coimbra, Portugal
4Faculty of Medicine, University of Coimbra, Coimbra, Portugal
- Direct-acting antivirals
- Hemodialysis patients
- Hepatitis C virus
- Kidney transplanted patients
- Occult hepatitis C infection
Abstract
Background: Occult Hepatitis C Infection (OCI) is characterized by the detection of Hepatitis C Virus (HCV) RNA in hepatocytes and in Peripheral Blood Mononuclear Cells (PBMCs) with no detection in serum, and PBMCs and Red Blood Cells (RBCs) were more recently suggested as predictors for OCI characterization. Here we aimed to evaluate OCI in Kidney Transplanted Patients (KTRs) and in Hemodialysis Patients (HD) treated with Direct-Acting Antivirals (DAAs) five to nine years after achieved a Sustained Virological Response (SVR).
Methods: Nineteen patients-SVR12 were included in this study, 12 in the KTRs and 7 in the HD study groups. HCV/OCI-RNA was screened in serum, plasma, PBMCs and RBCs five to nine years after SVR by droplet digital PCR (ddPCR) for OCI identification. Next-Generation Sequencing (NGS) was performed to characterize the viruses detected by ddPCR.
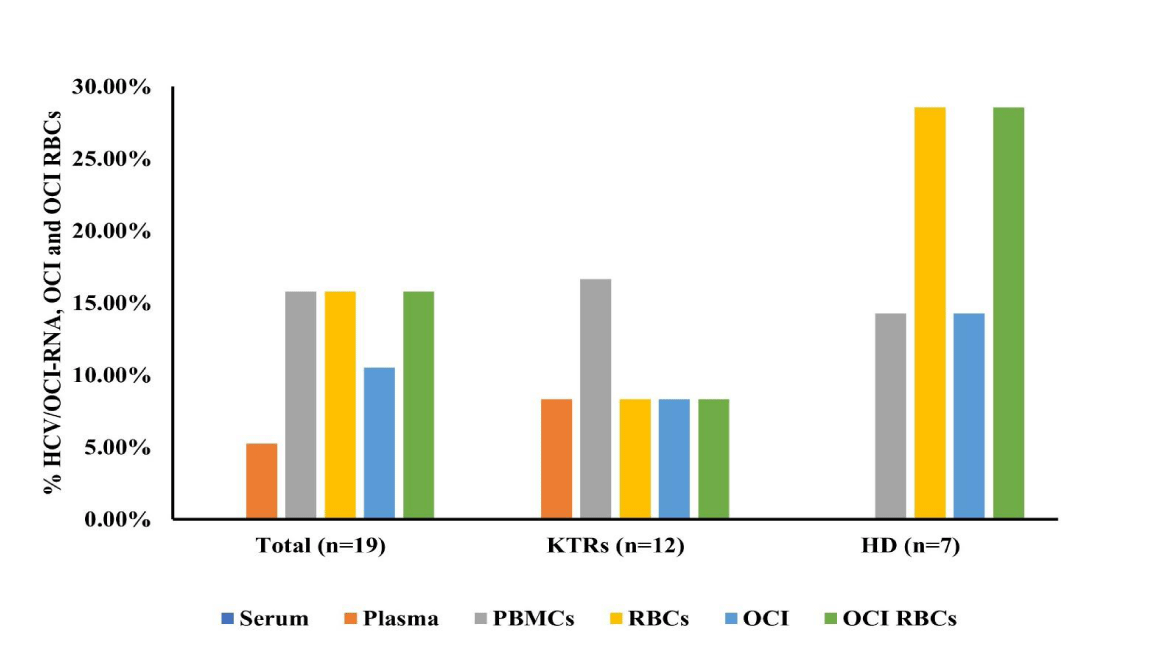
Results: In total, OCI (10.5%) and OCI RBCs (15.8%) patients were identified by ddPCR, 8.3% (both) of OCI (patient 7) and OCI RBCs (patient 9) in the KTRs, and 14.3% and 28.6% of OCI (patient 17) and OCI RBCs (patient 13 e 16) in the HD patients, respectively. HCV/OCI-RNA higher detection was in RBCs-HD (28.6%, n = 7) and PBMCs-KTRs (16.7%, n = 12) by ddPCR. HCV/OCI-RNA was not detected in analyzed samples by NGS.
Conclusion: OCI and OCI RBCs patients were identified in the KTRs and HD study groups by ddPCR suggesting that these patients might experience a relapse. PBMCs and RBCs could be predictors for HCV/OCI prevention, diagnosis and management in the study groups. These and a larger number of patients in both study groups should be evaluated in the future for a better understanding of these findings, as well as, their associated epidemiological and clinical significance.
Introduction
Hepatitis C Virus (HCV) infection is a substantial problem in some dialysis and kidney transplantation centres [1]. One of the high-risk groups for HCV infection are Haemodialysis patients (HD), and HCV infection was also suggested as a marker for graft loss and mortality among kidney transplanted patients (KTRs) [2,3]. Althought, Occult Hepatitis C Infection (OCI) that is defined by the detection of HCV Ribonucleic Acid (RNA) in hepatocytes and Peripheral Blood Mononuclear Cells (PBMCs) with no detection in serum [4], could be considered a concern for the transplantation centres and for viral transmission as is non-detected by currently screening methods [5]. Moreover, it could be considered as produced by chronic hepatitis C as no serological biomarkers of infection occur and there is not a large damage to the liver [6]. In this line there are several studies reporting OCI in HD patients worldwide, where mostly patients have anti-HCV antibodies and serum HCV-RNA serological markers of HCV infection repeatedly negative and abnormal liver enzyme levels [1, 6-16]. Furthermore, OCI was also reported in KTRs, including a study where the patients showed also repeatedly negative results for HCV serological biomarkers [1,7,14,17]. Nevertheless, in the literature no studies reporting OCI in HD and KTRs patients in Portugal were found.
Regarding Direct-Acting Antiviral Agents (DAAs) for HCV treatment in patients with Chronic Kidney Disease (CKD), there are studies describing DAAs efficacy with Sustained Virological Response (SVR) rounding 100%, including patients in haemodialysis, however, recent data, including patients on maintenance dialysis, call attention that many patients with advanced CKD could remain untreated, and also that numerous barriers to antiviral HCV treatment still exist [18-23]. There are also several reported KTRs studies, including a Portuguese study, describing high rates (e.g., 84%, > 91%, and 100%) of DAAs efficacy against HCV after SVR [24-29]. Additionally, a Portuguese study reporting two cases of hepatocellular carcinoma after DAAs therapy in KTRs infected with HCV was previously described [30]. Concerning OCI and DAAs, there are few studies in HD and KTRs patients, while, one study showed that OCI was identified in HD patients (5%, n = 60) who achieved a SVR after DAAs therapy, raising the concern that OCI could be present in these patients [12]. Moreover, Gelpi R, et al. [31,32]. Reported a study where OCI was excluded in treated (one treated with Peginterferon/Ribavirin and other with DAAs) donors for KTRs in 2018, and more recently, a OCI case in a patient with 3 kidney transplantations was described.
Recently, data regarding OCI in patients HCV negative by real-time PCR-baseline, in drug and non-drug users, including patients treated with DAAs that had achieved an SVR and with spontaneous clearance of HCV infection, and also the possibility of HCV/OCI transmission was described [5,33,34].
Here we aimed to evaluate OCI in anti-HCV positive HD and KTRs patients treated with DAAs five to nine years after achieved a SVR. For this, serum, plasma, PBMCs and Red Blood Cells (RBCs) samples in the both groups were screened for HCV/OCI-RNA detection by droplet digital PCR (ddPCR) as previously described [5,34], and OCI and OCI RBCs patients were identified as previously described [5, 34]. Additionally, Next-Generation Sequencing (NGS) was also performed in HCV/OCI-RNA positive samples of patients in the both study groups by ddPCR in order to characterize the detected viruses.
OCI and OCI RBCs patients were identified in this study and we hypothesized that these patients might experience a relapse.
Materials and Methods
Study groups
A total of 19 anti-HCV positive patients from the Coimbra Hospital and University Centre (CHUC) were included in this study in two study groups: KTRs - 12 kidney transplanted patients, and HD - 7 haemodialysis patients. All patients were previously treated with DAAs, at clinical evaluation time, and all achieved a SVR at week 12, by RT-PCR. Patients’ clinical parameters were collected and summed up in table 1. DAAs used for patients’ treatment were Sofosbuvir + Ledipasvir (SOF/LED), Ombitasvir + Paritaprevir + Ritonavir plus Dasabuvir (OMB/PAR/RIT + DAS), Glecaprevir + Pibrentasvir (GLE/PIB) and Elbasvir/Grazoprevir (ELB/GRZ), following the European Association for the Study of the Liver (EASL) recommendations. No patients positive and negative control groups were included in the study. Patients in the both study groups were evaluated for HCV/OCI-RNA five to nine years after treatment, during 2023 to 2024, by ddPCR. The CHUC Ethics Committee approved the study (registration number CHUC-146-19) and written informed consent was provided by all patients.
| Table 1: Clinical parameters of the patients in the KTRs and HD study groups at clinical evaluation time. | |||||||||||||||||||||
| Study Groups | Patients | Treatment | Sex | Age | Pre-KTRs Fbs | AST | ALT | GGT | INR | ALB | BT | #AH | #DM | #OB | #ALC | #TOB | #DRU | HCV- RNA Serum CET, RT-PCR | HCV genotype | DAAs | HCV- RNA Serum, RT-PCR baseline, SVR |
| KTRs | 1 | 2015 | F | 55 | + | - | - | - | - | - | - | + | - | - | - | - | - | + | 1a | SOF/LED | - |
| 2 | 2016 | M | 62 | - | + | + | - | - | - | - | + | - | + | - | - | - | + | 1b | SOF/LED | - | |
| 3 | 2015 | M | 68 | - | - | - | + | - | - | - | + | - | - | - | + | - | + | 1a | SOF/LED | - | |
| 4 | 2018 | M | 58 | - | + | + | + | - | - | - | + | - | - | - | - | - | + | 3a | GLE+PIB | - | |
| 5 | 2015 | M | 64 | - | + | + | + | + | - | - | + | - | + | - | - | - | + | 1b | SOF/LED | - | |
| 6 | 2015 | M | 55 | - | + | + | - | - | - | - | + | - | - | - | - | - | + | 1b | SOF/LED | - | |
| 7 | 2017 | F | 57 | - | + | - | - | - | - | - | + | - | - | - | - | - | + | 1b | SOF/LED | - | |
| 8 | 2015 | M | 51 | - | - | - | - | - | + | - | + | - | - | - | - | - | + | 1b | SOF/LED | - | |
| 9 | 2016 | M | 56 | - | - | - | - | + | - | - | - | - | - | - | - | - | + | 1b | SOF/LED | - | |
| 10 | 2015 | M | 64 | - | - | - | - | + | - | + | + | + | - | - | - | - | + | 1a | SOF/LED | - | |
| 11 | 2015 | M | 67 | - | - | - | - | - | - | - | + | + | - | - | + | - | + | 1b | OMB/PAR/RIT +DAS | - | |
| 12 | 2015 | M | 55 | + | - | - | - | - | - | - | + | - | - | - | - | - | + | 1a | SOF/LED | - | |
| HD | 13 | 2016 | M | 61 | - | - | - | - | - | - | - | + | - | - | + | - | - | + | 1b | OMB/PAR/RIT +DAS | - |
| 14 | 2016 | M | 61 | - | - | - | - | - | - | - | + | + | - | - | - | - | + | 1b | OMB/PAR/RIT +DAS | - | |
| 15 | 2018 | M | 64 | - | + | + | - | + | - | - | + | - | + | - | + | - | + | 3a | GLE+PIB | - | |
| 16 | 2018 | M | 79 | - | - | - | - | - | + | - | + | + | - | + | - | - | + | 1b | GLE+PIB | - | |
| 17 | 2016 | F | 42 | - | - | - | - | + | - | - | + | - | - | - | - | - | + | 1a | OMB/PAR/RIT +DAS | - | |
| 18 | 2016 | M | 67 | + | - | - | - | - | - | - | + | + | - | - | + | - | + | 1b | OMB/PAR/RIT +DAS | - | |
| 19 | 2017 | F | 71 | - | + | + | + | - | + | - | + | + | - | - | - | - | + | 1b | ELB/GRZ | - | |
| KTRs: Kidney Transplanted Recipients; HD- Haemodialysis; AST: Aspartate Aminotransferase; ALT: Alanine Aminotransferase; GTT: Gamma Glutamyl Transpeptidase; INR: International Normalized Ratio; ALB: Albumin; BT: Total Bilirubin; AH: Arterial Hypertension; DM- Diabetes; OB: Obesity; ALC: Alcohol; TOB: Tobacco; DRU: Drugs; HCV: Hepatitis C Virus; CET: Clinical Evaluation Time; RT-PCR, Quantitative Real-Time PCR; DAAs: Direct Antiviral Agents; SOF/LED: Sofosbuvir + Ledipasvir; OMB/PAR/RIT + DAS: Ombitasvir + Paritaprevir + Ritonavir Plus Dasabuvir; GLE/PIB: Glecaprevir + Pibrentasvir; ELB/GRZ: Elbasvir/Grazoprevir; SVR- Sustained Virological Response; OCI: Occult Hepatitis C Infection; F: Female; M: Male; *(-) Normal values: AST < 35 U/L; ALT < 45 U/L; GTT < 55 U/L; (+) Altered value. **Reference values: ALB 3.5-5.2 g/dL; Total BIL 0.2-1.2 mg/dL, INR 0.81-1.19; (-) Normal value; (+) Altered value. ***Reference values Fibroscan (Fbs): F0/F1: < 7 kPa; F2: 7-9.5 kPa; F3: 9.6-12.5; F3/F4: 12.5-14.5 kPa; F4: > 14.5 kPa; (-) absent fibrosis; (+) Fibrosis. # (-) Without, (+) With. (-) Negative result - RT-PCR, (+) Positive result - RT-PCR. | |||||||||||||||||||||
Samples collection
Blood (serum, plasma, PBMCs and RBCs) samples of all patients/groups were collected and treated as previously described [5,34,35]. Briefly, serum was recovered from dry tubes. Plasma, PBMCs and RBCs were isolated from blood samples collected in lithium heparin tubes that were separated using lymphoprep™ (Alere Technologies AS, Norway). All samples were stored at - 80ºC until use. After, HCV/OCI-RNA was screened in serum, plasma, PBMCs and RBCs resuspended in water molecular grade from all patients/groups by ddPCR, baseline. Additionally, NGS was also performed in HCV/OCI-RNA positive KTRs and HD patients’ analysed samples by ddPCR in order to characterize the detected viruses.
RNA extraction and cDNA synthesis
Total RNA was extracted from serum, plasma, PBMCs and RBCs resuspended in water molecular grade samples (each 250 μL) of all patients in the KTRs and HD study groups using TRI Reagent LS (Sigma-Aldrich, Germany) as previously described [5,33,34,36,37]. For cDNA synthesis the Xpert cDNA synthesis kit (GRISP, Portugal) was used following the manufactures instructions as previously described [5,34].
ddPCR: HCV/OCI-RNA was detected by ddPCR for the HCV core region using sense/antisense (5′-GCGTTAGTAYGAGTGTYG/5′-CRATTCCGGTGTACTCAC) primers, and the FAM-labeled HCV probe 5′-FAM-CCGCAGACCACTATGGCTC-BHQ1–3′ as previously described [5,34]. The plasmid p7-nsGluc2A (pGNN-nonreplicative), kindly provided by Professor Charles Rice (Apath, L.L.C and Rockefeller University, USA), was used as positive control in the ddPCR reactions as previously described [5]. Here, the samples limit of HCV/OCI-RNA detection was of 0.39 copies/μL (lower, RBCs) and 0.83 copies/μL (higher, RBCs), and for the positive control (plasmid) was of 332 copies/μL (lower) and 2.992 copies/μL (higher), all with accepted droplets greater than 10.000. OCI patients’ identification was done considering ddPCR PBMCs positive and serum negative results as previously described [5,34]. OCI RBCs patients’ identification was done considering ddPCR RBCs positive and serum negative results as previously described [5,34].
NGS: After RNA extraction of the HCV/OCI-RNA positive samples by ddPCR, plasma of patient 1 and PBMCs of the patients 1 and 7 (OCI) in the KTRs group, and RBCs of patients 13 and 16 in the HD group, both OCI RBCs, were sequenced by NGS following previously described data with some modifications [38-40]. Briefly, relatively to Di H, et al. [40]. Previously reported data here turbo DNase kit (Thermo Fisher Scientific, USA) was used for RNA treatment following the manufactures instructions, and RNeasy Mini Kit (Qiagen, Germany) was used to clean the RNA products following the manufacturer’s instructions. RNA libraries were performed as previously described [38,39]. Briefly, human rRNA was removed using the NEBNext rRNA depletion kit (New England Biolabs, USA), and RNA libraries were prepared with the NEBNext Ultra II directional RNA library preparation kit (New England Biolabs, USA) following the manufactures instructions [38,39]. For each sample the final RNA library was sequenced using paired-end 150-bp reads on an Illumina Novaseq6000 instrument (Illumina Inc., USA) [38,39] at Macrogen (Macrogen Inc, South Korea). After, the resulting pair-end-reads were trimmed using Trimmomatic [41] and the host genome sequences were removed using Bowtie2 [42], with the RefSeq human genome (GCF_000001405.40_ GRCh38.p14) as a reference, that is available at the National Center for Biotechnology Information (NCBI). Then, the remaining reads were de novo assembled using Megahit V1.2.9 [43], and the de novo assembled contigs were, in a first step, screened for putative viral matches against the NCBI RefSeq Virus database using Diamond [44]. To remove false positive hits, the putative viral contigs (> 500 nucleotides in length and an e-value ≤ 105) were further tested with NCBI BLASTx/BLASTn searches against the refseq protein and nucleotide (nr/nt) databases, respectively. HCV/OCI-RNA RBCs (patient 9-KTRs) and PBMCs (patient 17-HD) positive samples by ddPCR were not sequenced by NGS because they were recently collected and have shown a viral load (low) similarly to the sequenced ones (without HCV/OCI identified by NGS). All HCV/OCI-RNA positive samples by ddPCR will be inoculated in MDBK cell cultures as previously described [5] in order to characterize HCV/OCI-RNA in these samples.
Statistical analysis
Frequencies, percentages, and means were used for descriptive analysis in study group samples using Microsoft® Excel® 2016 MSO. HCV/OCI-RNA detected per well in copies/μL was validated by the Poisson statistics included in the Bio-Rad QuantaSoft™ Analysis Pro-Software v. 1.0.596 following the manufactures instructions, with the error bars representing Poisson 95% confidence intervals [34].
Results
ddPCR
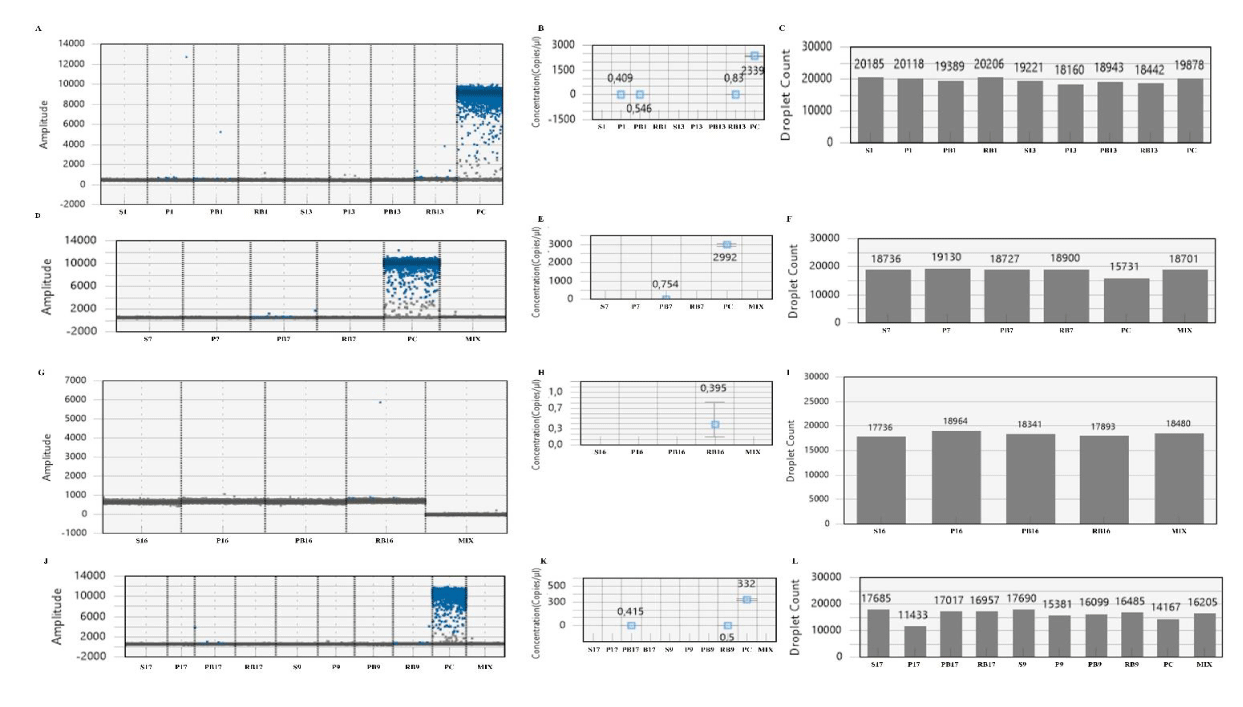
OCI (10.5%, n = 19) and OCI RBCs (15.8%, n = 19) were presented in the total study population by ddPCR (considering PBMCs positive and serum negative results for OCI, and RBCs positive and serum negative results for OCI RBCs classifications), corresponding to 8.3% (n = 12) (both) of OCI (patient 7) and OCI RBCs (patient 9) in the KTRs, and 14.3% (n = 7) and 28.6% (n = 7) of OCI (patient 17) and OCI RBCs (patients 13 and 16), respectively, in the HD study group (Figure 1). In total analysed blood samples, HCV/OCI-RNA was detected in 15.8% (n = 19) of the PBMCs and RBCs samples, and in 5.3% (n = 19) of the plasma samples by ddPCR (Figure 1). Within study groups, 16.7% (n = 12) and 8.3% (both, n = 12) were detected in the PBMCs, and in the RBCs and plasma of the patients in the KTRs group, respectively, and 28.6% (n = 7) and 14.3% (n = 7) were detected in RBCs and PBMCs, respectively, of the patients in the HD group. No HCV/OCI-RNA was detected in the total (n = 19) analysed serum samples in the both study groups (Figure 1). The viral load detected in the HCV/OCI-RNA positive analysed samples by ddPCR were 0.41 and 0.55 copies/μL in the plasma and PBMCs of the patient 1, 0.75 copies/μL in the PBMCs of the patient 7 (OCI) and 0.50 copies/μL in the RBCs of the patient 9 (OCI RBCs) in the KTRs group. Relatively to the HD study group, 0.83 and 0.39 copies/μL were detected in the RBCs of the patient 8 and 11 (OCI RBCs), respectively, and 0.41 copies/μL in the PBMCs of the patient 17 (OCI) (Figure 2 and table 2). DdPCR negative results obtained for the remaining tested patients’ samples/groups are shown in table 2. Blood samples fluorescence 1D amplitude plots for the patients with HCV/OCI-RNA positive results are shown in figure 2. In the KTRs group, patient 1 was positive for the plasma and PBMCs samples, and patient 7 was positive for the PBMCs sample and negative for the serum, plasma and RBCs samples (Figure 2). Patient 8 and 11 in the HD group, were both positive for the RBCs samples and negative for the serum, plasma and PBMCs samples (Figure 2). The lower and the higher limits of HCV/OCI-RNA detected in this study were 0.39 (RBCs) and 0.83 (RBCs) copies/μL, respectively (Figure 2 and table 2). The higher and the lower concentrations detected for the positive control (plasmid) were 2.992 and 332 copies/μL (Figure 2). All results were validated by the encountered number of positive and negative droplets relatively to the total obtained number of accepted droplets that were greater than 10.000 (Figure 2 and table 2).
| Table 2: DdPCR of analysed blood samples of the patients in the KTRs and HD study groups performed five to nine years after SVR. HCV/OCI-RNA and droplets count detected, and OCI and OCI RBCs patients’ identification. Clinical parameters of the patients in the KTRs and HD study groups are shown in table 2. | ||||||||
| Study Groups | Year of patients treatment | Year of ddPCR runs | Patient Sample | HCV/OCI-RNA (copies/µL) | Accepted droplets | Positive droplets | Negative droplets | *OCI or OCI RBCs |
| KTRs | 2015 | 2023 | 1 - Serum | 0.00 | 20185 | 0 | 20185 | |
| 1 - Plasma | 0.41 | 20118 | 7 | 20111 | ||||
| 1 - PBMCs | 0.55 | 19389 | 9 | 19380 | ||||
| 1 - RBCs | 0.00 | 20206 | 0 | 20206 | ||||
| KTRs | 2016 | 2023 | 2 - Serum | 0.00 | 19887 | 0 | 19887 | |
| 2 - Plasma | 0.00 | 19992 | 0 | 19992 | ||||
| 2 - PBMCs | 0.00 | 18791 | 0 | 18791 | ||||
| 2 - RBCs | 0.00 | 19827 | 0 | 19827 | ||||
| KTRs | 2015 | 2023 | 3 - Serum | 0.00 | 18941 | 0 | 18941 | |
| 3 - Plasma | 0.00 | 19579 | 0 | 19579 | ||||
| 3 - PBMCs | 0.00 | 19336 | 0 | 19336 | ||||
| 3 - RBCs | 0.00 | 19774 | 0 | 19774 | ||||
| KTRs | 2018 | 2023 | 4 - Serum | 0.00 | 17787 | 0 | 17787 | |
| 4 - Plasma | 0.00 | 19126 | 0 | 19126 | ||||
| 4 - PBMCs | 0.00 | 19815 | 0 | 19815 | ||||
| 4 - RBCs | 0.00 | 19854 | 0 | 19854 | ||||
| KTRs | 2015 | 2023 | 5 - Serum | 0.00 | 18517 | 0 | 18517 | |
| 5 - Plasma | 0.00 | 19373 | 0 | 19373 | ||||
| 5 - PBMCs | 0.00 | 18617 | 0 | 18617 | ||||
| 5 - RBCs | 0.00 | 18208 | 0 | 18208 | ||||
| KTRs | 2015 | 2023 | 6 - Serum | 0.00 | 19389 | 0 | 19389 | |
| 6 - Plasma | 0.00 | 17984 | 0 | 17984 | ||||
| 6 - PBMCs | 0.00 | 18490 | 0 | 18490 | ||||
| 6 - RBCs | 0.00 | 18286 | 0 | 18286 | ||||
| KTRs | 2017 | 2023 | 7 - Serum | 0.00 | 18736 | 0 | 18736 | OCI |
| 7 - Plasma | 0.00 | 19130 | 0 | 19130 | ||||
| 7 - PBMCs | 0.75 | 18727 | 12 | 18715 | ||||
| 7 - RBCs | 0.00 | 18900 | 0 | 18900 | ||||
| KTRs | 2015 | 2024 | 8 - Serum | 0.00 | 17428 | 0.00 | 17428 | |
| 8 - Plasma | 0.00 | 18714 | 0.00 | 18714 | ||||
| 8 - PBMCs | 0.00 | 10094 | 0.00 | 10094 | ||||
| 8 - RBCs | 0.00 | 17985 | 0.00 | 17985 | ||||
| KTRs | 2016 | 2024 | 9 - Serum | 0.00 | 17690 | 0.00 | 17690 | OCI RBCs |
| 9 - Plasma | 0.00 | 15381 | 0.00 | 15381 | ||||
| 9 - PBMCs | 0.00 | 16099 | 0.00 | 16099 | ||||
| 9 - RBCs | 0.50 | 16485 | 7.00 | 16478 | ||||
| KTRs | 2015 | 2024 | 10 - Serum | 0.00 | 18162 | 0.00 | 18162 | |
| 10 - Plasma | 0.00 | 11267 | 0.00 | 11267 | ||||
| 10 - PBMCs | 0.00 | 16302 | 0.00 | 16302 | ||||
| 10 - RBCs | 0.00 | 17920 | 0.00 | 17920 | ||||
| KTRs | 2015 | 2024 | 11 - Serum | 0.00 | 18428 | 0.00 | 18428 | |
| 11 - Plasma | 0.00 | 16197 | 0.00 | 16197 | ||||
| 11 - PBMCs | 0.00 | 16151 | 0.00 | 16151 | ||||
| 11 - RBCs | 0.00 | 18361 | 0.00 | 18361 | ||||
| KTRs | 2015 | 2024 | 12 - Serum | 0.00 | 18550 | 0.00 | 18550 | |
| 12 - Plasma | 0.00 | 18525 | 0.00 | 18525 | ||||
| 12 - PBMCs | 0.00 | 16195 | 0.00 | 16195 | ||||
| 12 - RBCs | 0.00 | 17071 | 0.00 | 17071 | ||||
| HD | 2016 | 2023 | 13 - Serum | 0.00 | 19221 | 0 | 19221 | OCI RBCs |
| 13 - Plasma | 0.00 | 20459 | 0 | 20459 | ||||
| 13 - PBMCs | 0.00 | 18943 | 0 | 18943 | ||||
| 13 - RBCs | 0.83 | 18442 | 13 | 18429 | ||||
| HD | 2016 | 2023 | 14 - Serum | 0.00 | 18318 | 0 | 18318 | |
| 14 - Plasma | 0.00 | 19186 | 0 | 19186 | ||||
| 14 - PBMCs | 0.00 | 18301 | 0 | 18301 | ||||
| 14 - RBCs | 0.00 | 19167 | 0 | 19167 | ||||
| HD | 2018 | 2023 | 15 - Serum | 0.00 | 18285 | 0 | 18285 | |
| 15 - Plasma | 0.00 | 19575 | 0 | 19575 | ||||
| 15 - PBMCs | 0.00 | 19223 | 0 | 19223 | ||||
| 15 - RBCs | 0.00 | 19161 | 0 | 19161 | ||||
| HD | 2018 | 2023 | 16 - Serum | 0.00 | 17736 | 0 | 17736 | OCI RBCs |
| 16 - Plasma | 0.00 | 18964 | 0 | 18964 | ||||
| 16 - PBMCs | 0.00 | 18440 | 0 | 18440 | ||||
| 16 - RBCs | 0.39 | 17893 | 6 | 17887 | ||||
| HD | 2016 | 2024 | 17 - Serum | 0.00 | 17685 | 0.00 | 17685 | OCI |
| 17 - Plasma | 0.00 | 11433 | 0.00 | 11433 | ||||
| 17 - PBMCs | 0.41 | 17017 | 6.00 | 17011 | ||||
| 17 - RBCs | 0.00 | 16957 | 0.00 | 16957 | ||||
| HD | 2016 | 2024 | 18 - Serum | 0.00 | 17334 | 0.00 | 17334 | |
| 18 - Plasma | 0.00 | 17391 | 0.00 | 17391 | ||||
| 18 - PBMCs | 0.00 | 16600 | 0.00 | 16600 | ||||
| 18 - RBCs | 0.00 | 17613 | 0.00 | 17613 | ||||
| HD | 2017 | 2024 | 19 - Serum | 0.00 | 18240 | 0.00 | 18240 | |
| 19 - Plasma | 0.00 | 14883 | 0.00 | 14883 | ||||
| 19 - PBMCs | 0.00 | 16385 | 0.00 | 16385 | ||||
| 19 - RBCs | 0.00 | 15154 | 0.00 | 15154 | ||||
| *OCI: Serum negative; PBMCs positive; OCI RBCs: Serum negative; RBCs positive. | ||||||||
NGS
No putative contig related to HCV was found in sequenced samples (n = 5) by NGS, plasma of patient 1 and PBMCs of patients 1 and 7 (OCI patient) in the KTRs (n = 3) group, and RBCs of patients 13 and 16 in the HD (n = 2) group, both OCI RBCs, all with HCV/OCI-RNA detected by ddPCR.
Discussion
HCV infection is a substantial problem in dialysis and kidney transplantation centres, being HD patients considered one high-risk group for this infection and HCV infection suggested as a risk factor for graft loss and mortality among KTRs patients [1-3]. Moreover, higher prevalence rate of HCV infection was reported in the dialysis population relatively to patients without kidneys clinical complications, and this occurs in either developing or developed countries [45,46]. Furthermore, currently, nosocomial transmission of HCV was linked to HCV spread within HD units all over the world [46]. Concerning KTRs, the prevalence of HCV infection varies according to factors, such as, the prevalence of HCV infection in dialysis units, the type of dialysis, and the duration of the dialytic therapy, and varies from 5% to 50% in the developed world [47]. Althought, OCI, that is characterized by the detection of HCV-RNA in hepatocytes and PBMCs without detection in serum by current screening methods, may be under-recognized in HD and KTRs patients causing concern in the possibility of patients that tested negatively for HCV continued to be infected and also in the spread of the virus [4,5,13]. There are several studies reporting OCI worldwide, mainly in HD but also in KTRs patients, causing also concern, as the majority of them described the OCI identification in patients that have anti-HCV antibodies and serum HCV-RNA serological markers of HCV infection repeatedly negative, irrespective of the level of the liver enzymes [1,6,17]. Nevertheless, so far, there are no studies in the literature reporting OCI in HD or KTRs patients in Portugal.
Concerning DAAs for HCV treatment, there are studies describing high rates of DAAs efficacy after SVR (e.g., 84%, > 91%, and 100%) in patients with CKD including HD and KTRs patients [18-21,24-29]. However, recently Fabrizi F, et al. [20]. Referred that reported data highlight to the possibility of numerous patients with advanced CKD could not be treated, that remains numerous barriers to HCV antiviral treatment, and that is also needed to overcome the current barriers for HCV treatment in dialysis patients. Moreover, a Portuguese study reporting two cases of hepatocellular carcinoma after DAAs therapy in KTRs infected with HCV was previously described [30].
Here, we aimed to evaluate OCI in anti-HCV positive HD and KTRs patients treated with DAAs five to nine years after achieved a SVR.
OCI (10.5%, n = 19) and OCI RBCs (15.8%, n = 19) were identified in the total analysed patients’ samples of the both study groups by ddPCR. OCI was identified in 1 KTRs (8.3%, n = 12) and in 1 HD (14.3%, n = 7) patients, and OCI RBCs was identified in 1 KTRs (8.3%, n = 12) and in 2 HD (28.6%, n = 7) patients. Moreover, patient 1 in the KTRs group was also plasma and PBMCs HCV/OCI-RNA positive by ddPCR. We have found few studies in HD and KTRs patients who achieved an SVR after DAAs therapy, however, recently, Naguib H, et al. [12]. Reported the first study in HD patients treated with DAAs, SVR 24, and have identified 3 (n = 60) OCI patients in line with our results. The question about OCI-concern was aborded again and we corroborate this question. In 2018 Gelpi R, et al. [31]. Reported a study where OCI was excluded in a donor for KTRs treated with DAAs and recently OCI was described in a study involving KTRs (n = 51), where OCI was identified in 1 patient [32], similar to our results. Overall results suggest that OCI/OCI RBCs can be detected within HD and KTRs patients with apparent clearance of HCV-RNA in serum and arguing the questions if these patients might experience a relapse or are cured, and there are studies suggesting OCI as a higher risk factor for relapse, disease progression and HCV transmission to others [5,12,23]. The persistence of detection of low levels of HCV-RNA in PBMCs and/or liver, without detection of viremia in serum, SVR after DAAs therapy-baseline or SVR after DAAs therapy-follow-up, including patients with liver transplantation, that could lead to important consequences for patient’s clinical outcomes were also previously described [14,48,49]. Additionally, there are studies reporting the detection of HCV by NGS, including patients with DAAs therapy [38-51], while we did not found studies reporting OCI by NGS in the literature and our results are accordantly. The presence of the low viral load detected in the analysed samples by ddPCR could be an explanation of the achieved results, and similar OCI viral loads were previously reported, abording OCI in HCV patients who achieved an SVR after DAAs therapy [5,34]. Moreover, ddPCR [52]. was considered as a valuable addition to virologist`s due to main features, such as, high specificity, sensitivity and reproducibility, no standard curve is needed for absolute quantification, and high efficacy when compared to conventional molecular methods [53]. Here, OCI and OCI RBCs patients were identified in the KTRs and HD study groups by ddPCR suggesting that these patients might experience a relapse.
Conclusion
OCI was identified in 1 patient of the KTRs and HD study groups, and OCI RBCs was identified in 1 and 2 patients of the KTRs and HD study groups, respectively, by ddPCR. HCV/OCI-RNA was also identified in the plasma and PBMCs of another patient in the KTRs group by ddPCR. These patients might experience a relapse. HCV/OCI-RNA higher detection was in PBMCs (16.7%, n = 12) and RBCs (28.6%, n = 7) of the patients in the KTRs and HD study groups, respectively. DdPCR demonstrated a higher sensitivity to detect HCV/OCI-RNA compared to NGS. PBMCs and RBCs samples could be suggested as predictors for HCV/OCI prevention, diagnosis and management in KTRs and HD patients. To better understand these findings and their virological and clinical significance, studies involving follow-up of the studied patients, and a larger number of HD and KTRs patients should be performed in the future.
Acknowledgment
The authors acknowledge GILEAD GÉNESE under the grant PGG/004/2017. The authors acknowledge Raquel Monteiro from the CHUC for her help in the blood samples collection. The authors also acknowledge the Cancer Biology & Epigenetics Group, Research Centre of IPO Porto, the Molecular Pathology and Immunogenetic Laboratories, ICBAS-UP, for their availability to carry out the work.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Dolatimehr F, Khosravi MH, Rezaee-Zavareh MS, Alavian SM. Prevalence of occult HCV infection in emodialysis and kidney-transplanted patients: A systematic review. Future Virology. 2017;12(5):315-22. doi: 10.2217/fvl-2016-0138.
- Rostami Z, Nourbala MH, Alavian SM, Bieraghdar F, Jahani Y, Einollahi B. The impact of Hepatitis C virus infection on kidney transplantation outcomes: A systematic review of 18 observational studies: The impact of HCV on renal transplantation. Hepat Mon. 2011;11(4):247-54. PMID: 22087151.
- Alavian SM, Kabir A, Ahmadi AB, Lankarani KB, Shahbabaie MA, Ahmadzad-Asl M. Hepatitis C infection in hemodialysis patients in Iran: a systematic review. Hemodial Int. 2010;14(3):253-62. doi: 10.1111/j.1542-4758.2010.00437.x. PMID: 20491973.
- Castillo I, Pardo M, Bartolomé J, Ortiz-Movilla N, Rodríguez-Iñigo E, de Lucas S, Salas C, Jiménez-Heffernan JA, Pérez-Mota A, Graus J, López-Alcorocho JM, Carreño V. Occult hepatitis C virus infection in patients in whom the etiology of persistently abnormal results of liver-function tests is unknown. J Infect Dis. 2004;189(1):7-14. doi: 10.1086/380202. PMID: 14702147.
- Silva E, Marques S,Osorio H, Canhão B, Madaleno J, Simão A, Carvalho A. Occult hepatitis C infection: Viruses with infectious potential in Huh7.5 and MDBK cell lines suggest HCV/OCI transmission. Pharmacy & Pharmacology International Journal. 2023. doi: 10.15406/ppij.2023.11.00394.
- El-Shishtawy S, Sherif N, Abdallh E, Kamel L, Shemis M, Saleem AA, Abdalla H, El Din HG. Occult Hepatitis C Virus Infection in Hemodialysis Patients; Single Center Study. Electron Physician. 2015;7(8):1619-25. doi: 10.19082/1619. PMID: 26816589.
- Baid-Agrawal S, Schindler R, Reinke P, Staedtler A, Rimpler S, Malik B, Frei U, Berg T. Prevalence of occult hepatitis C infection in chronic hemodialysis and kidney transplant patients. Journal of Hepatology. 2014;60(5):928-33. doi: 10.1016/j.jhep.2014.01.012.
- Bayani M, Hasanjani Roushan MR, Soleimani Amiri MJ, Yahyapour Y, Ebrahimpour S, Akbarzadeh Jelodar S. Prevalence of occult hepatitis C virus infection in hemodialysis patients. Caspian J Intern Med. 2021;12(4):533-8. doi: 10.22088/cjim.12.4.533. PMID: 34820059.
- Naghdi R, Ranjbar M, Bokharaei-Salim F, Keyvani H, Savaj S, Ossareh S, Shirali A, Mohammad-Alizadeh AH. Occult Hepatitis C Infection Among Hemodialysis Patients: A Prevalence Study. Ann Hepatol. 2017;16(4):510-3. doi: 10.5604/01.3001.0010.0277. PMID: 28611267.
- Abdelmoemen G, Khodeir SA, Abou-Saif S, Kobtan A, Abd-Elsalam S. Prevalence of occult hepatitis C virus among hemodialysis patients in Tanta university hospitals: a single-center study. Environ Sci Pollut Res Int. 2018;25(6):5459-64. doi: 10.1007/s11356-017-0897-y. PMID: 29214477.
- Nakhaie M, Taheri E, Charostad J, Arefinia N, Kalantar-Neyestanaki D, Rezaei Zadeh Rukerd M, Ahmadpour F, Pourebrahimi MH, Ahmadinejad Farsangi S, Shafieipour S. Prevalence of hepatitis C virus and its occult infection in hemodialysis patients. Jundishapur J Microbiol. 2023;16(5):e136504. doi: 10.5812/jjm-136504.
- Naguib H, Abouelnaga SF, Elsayed MM. Occult hepatitis C virus infection in hemodialysis patients who achieved a sustained virological response to directly acting antiviral drugs: is it a concern? Int Urol Nephrol. 2024;56(1):217-22. doi: 10.1007/s11255-023-03621-1. PMID: 37209238.
- Austria A, Wu GY. Occult Hepatitis C Virus Infection: A Review. J Clin Transl Hepatol. 2018;6(2):155-60. doi: 10.14218/jcth.2017.00053. PMID: 29951360.
- Wróblewska A, Bielawski KP, Sikorska K. Occult Infection with hepatitis C virus: Looking for clear-cut boundaries and methodological consensus. Journal of Clinical Medicine. 2021;10(24):5874. doi: 10.3390/jcm10245874.
- Taherpour S, Javanmard D, Ziaee M. Occult Hepatitis C Virus Infection among Hemodialysis Patients: An Iranian Experience. Arch Iran Med. 2020;23(9):586-92. Epub 20200901. doi: 10.34172/aim.2020.68. PMID: 32979904.
- Ali NK, Mohamed RR, Saleh BE, Alkady MM, Farag ES. Occult hepatitis C virus infection among haemodialysis patients. Arab J Gastroenterol. 2018;19(3):101-5. Epub 20180920. doi: 10.1016/j.ajg.2018.09.001. PMID: 30245116.
- Jiménez C, Olea T, Santana M, Lopez M, Castillo I, Bartolomé J, Quiroga J, Vaca M, Carreño V, Selgas R. Occult Hepatitis C Virus in Recipients of Kidney Transplantation: Prevalence and Clinical Implications. Transplantation. 2018;102:S658. doi: 10.1097/01.tp.0000543589.88342.67. PMID: 00007890-201807001-01058.
- Iliescu EL, Mercan-Stanciu A, Toma L. Safety and efficacy of direct-acting antivirals for chronic hepatitis C in patients with chronic kidney disease. BMC Nephrology. 2020;21(1):21. doi: 10.1186/s12882-020-1687-1.
- Agarwal SK, Bagchi S, Yadav RK. Hemodialysis Patients Treated for Hepatitis C Using a Sofosbuvir-based Regimen. Kidney Int Rep. 2017;2(5):831-5. doi: 10.1016/j.ekir.2017.04.003. PMID: 29270489.
- Fabrizi F, Cerutti R, Messa P. An updated view on the antiviral therapy of hepatitis C in chronic kidney disease. Pathogens. 2021;10(11):1381. doi: 10.3390/pathogens10111381.
- Ryu JE, Song MJ, Kim SH, Kwon JH, Yoo SH, Nam SW, Nam HC, Kim HY, Kim CW, Yang H, Bae SH, Song DS, Chang UI, Yang JM, Lee SW, Lee HL, Lee SK, Sung PS, Jang JW, Choi JY, Yoon SK. Safety and effectiveness of direct-acting antivirals in patients with chronic hepatitis C and chronic kidney disease. Korean J Intern Med. 2022;37(5):958-68. doi: 10.3904/kjim.2021.486. PMID: 35981893.
- Yousif MM, Elsadek Fakhr A, Morad EA, Kelani H, Hamed EF, Elsadek HM, Zahran MH, Fahmy Afify A, Ismail WA, Elagrody AI, Ibrahim NF, Amer FA, Zaki AM, Sadek A, Shendi AM, Emad G, Farrag HA. Prevalence of occult hepatitis C virus infection in patients who achieved sustained virologic response to direct-acting antiviral agents. Infez Med. 2018;26(3):237-43. PMID: 30246766.
- Mekky MA, Sayed HI, Abdelmalek MO, Saleh MA, Osman OA, Osman HA, Morsy KH, Hetta HF. Prevalence and predictors of occult hepatitis C virus infection among Egyptian patients who achieved sustained virologic response to sofosbuvir/daclatasvir therapy: a multi-center study. Infect Drug Resist. 2019;12:273-9. doi: 10.2147/idr.S181638. PMID: 30774394.
- Kamar N, Marion O, Rostaing L, Cointault O, Ribes D, Lavayssière L, Esposito L, Del Bello A, Métivier S, Barange K, Izopet J, Alric L. Efficacy and Safety of Sofosbuvir-Based Antiviral Therapy to Treat Hepatitis C Virus Infection After Kidney Transplantation. Am J Transplant. 2016;16(5):1474-9. doi: 10.1111/ajt.13518. PMID: 26587971.
- Colombo M, Aghemo A, Liu H, Zhang J, Dvory-Sobol H, Hyland R, Yun C, Massetto B, Brainard DM, McHutchison JG, Bourlière M, Peck-Radosavljevic M, Manns M, Pol S. Treatment With Ledipasvir-Sofosbuvir for 12 or 24 Weeks in Kidney Transplant Recipients With Chronic Hepatitis C Virus Genotype 1 or 4 Infection: A Randomized Trial. Ann Intern Med. 2017;166(2):109-17. doi: 10.7326/m16-1205. PMID: 27842383.
- Morales AL, Liriano-Ward L, Tierney A, Sang M, Lalos A, Hassan M, Nair V, Schiano T, Satoskar R, Smith C. Ledipasvir/sofosbuvir is effective and well tolerated in postkidney transplant patients with chronic hepatitis C virus. Clin Transplant. 2017;31(5). doi: 10.1111/ctr.12941. PMID: 28239909.
- Reau N, Kwo PY, Rhee S, Brown RS, Agarwal K, Angus P, Gane E, Kao JH, Mantry PS, Reddy KR, Tran TT, Hu YB, Gulati A, Ng TI, Dumas EO, Shulman N, Trinh R, Forns X. LBO-03 - MAGELLAN-2: Safety and efficacy of glecaprevir/pibrentasvir in liver or renal transplant adults with chronic hepatitis C genotype 1–6 infection. Journal of Hepatology. 2017;66(1, Supplement):S90-S1. doi: 10.1016/S0168-8278(17)30444-0.
- Chute DF, Chung RT, Sise ME. Direct-acting antiviral therapy for hepatitis C virus infection in the kidney transplant recipient. Kidney International. 2018;93(3):560-7. doi: 10.1016/j.kint.2017.10.024.
- Weigert A, Querido S, Carvalho L, Lebre L, Chagas C, Matias P, Birne R, Nascimento C, Jorge C, Adragão T, Bruges M, Machado D. Hepatitis C Virus Eradication in Kidney Transplant Recipients: A Single-Center Experience in Portugal. Transplant Proc. 2018;50(3):743-5. doi: 10.1016/j.transproceed.2018.02.017. PMID: 29661427.
- Calça R, Jorge C, Lebre L, Cacheira E, Querido S, Nascimento C, Adragão T, Bruges M, Weigert A, Machado D. Hepatocellular carcinoma after direct-acting antiviral therapy in kidney transplant recipients infected with hepatitis C virus. Nefrología (English Edition). 2020;40(6):675-6. doi: 10.1016/j.nefroe.2020.11.006.
- Gelpi R, Rodríguez-Villar C, Paredes D, Roque R, Ruiz A, Adalia R, de Sousa E, Revuelta I, Sole M, Martinez M, Navarro M, Lens S, Oppenheimer F, Diekmann F. Safety of hepatitis C virus (HCV)-treated donors for kidney transplantation excluding occult HCV infection through kidney biopsies. Transplant International. 2018;31(8):938-9. doi: 10.1111/tri.13270.
- Czarnecka P, Czarnecka K, Tronina O, Bączkowska T, Wyczałkowska-Tomasik A, Durlik M, Czerwinska K. Evaluation of long-term outcomes of direct acting antiviral agents in chronic kidney disease subjects: A single center cohort study. J Clin Med. 2023;12(10):3513. doi: 10.3390/jcm12103513.
- Silva E, Marques S, Salta S, Serqueira JP, Madaleno J, Simão A, Carvalho A. Occult hepatitis C infection detection in people who use drugs with or without direct-antiviral agents therapy. J Hepatol. 2022.
- Silva E, Marques S, Leal B, Canhao B, Madaleno J, Simao A, Carvalho A. Occult hepatitis C infection identified in injection drug users with direct antiviral agents therapy and spontaneous resolution of hepatitis C virus infection. Virus Res. 2023;329:199104. doi: 10.1016/j.virusres.2023.199104. PMID: 37003528.
- Costa-Matos L, Batista P, Monteiro N, Henriques P, Girao F, Carvalho A. Hfe mutations and iron overload in patients with alcoholic liver disease. Arq Gastroenterol. 2013;50(1):35-41. doi: 10.1590/s0004-28032013000100008. PMID: 23657305.
- Silva E, Marques S, Osório H, Carvalheira J, Thompson G. Endogenous hepatitis C virus homolog fragments in European rabbit and hare genomes replicate in cell culture. PLoS One. 2012;7(11):e49820. doi: 10.1371/journal.pone.0049820. PMID: 23185448.
- Silva E, Osório H, Thompson G. Hepatitis C-like viruses are produced in cells from rabbit and hare DNA. Sci Rep. 2015;5:14535. doi: 10.1038/srep14535. PMID: 26416202.
- Pedersen MS, Mollerup S, Nielsen LG, Jenssen H, Bukh J, Schønning K. Genome Sequence of an Unknown Subtype of Hepatitis C Virus Genotype 6: Another Piece for the Taxonomic Puzzle. Microbiol Resour Announc. 2019;8(42). doi: 10.1128/mra.01030-19. PMID: 31624172.
- Pedersen MS, Fahnøe U, Madsen LW, Christensen PB, Øvrehus A, Bukh J. Characterization of a Novel Hepatitis C Virus Genotype 1 Subtype from a Patient Failing 4 Weeks of Glecaprevir-Pibrentasvir Treatment. Microbiol Resour Announc. 2021;10(41):e0075521. doi: 10.1128/mra.00755-21. PMID: 34647796.
- Di H, Thor SW, Trujillo AA, Stark TJ, Marinova-Petkova A, Jones J, Wentworth DE, Barnes JR, Davis CT. Comparison of nucleic acid extraction methods for next-generation sequencing of avian influenza A virus from ferret respiratory samples. Journal of Virological Methods. 2019;270:95-105. doi: 10.1016/j.jviromet.2019.04.014.
- Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114-20. doi: 10.1093/bioinformatics/btu170. PMID: 24695404.
- Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9(4):357-9. doi: 10.1038/nmeth.1923. PMID: 22388286; PubMed Central PMCID: PMC3322381.
- Li D, Luo R, Liu CM, Leung CM, Ting HF, Sadakane K, Yamashita H, Lam TW. MEGAHIT v1.0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods. 2016;102:3-11. doi: 10.1016/j.ymeth.2016.02.020. PMID: 27012178.
- Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nature Methods. 2015;12(1):59-60. doi: 10.1038/nmeth.3176.
- Jadoul M, Bieber BA, Martin P, Akiba T, Nwankwo C, Arduino JM, Goodkin DA, Pisoni RL. Prevalence, incidence, and risk factors for hepatitis C virus infection in hemodialysis patients. Kidney Int. 2019;95(4):939-47. doi: 10.1016/j.kint.2018.11.038. PMID: 30904068.
- Fabrizi F, Cerutti R, Messa P. Updated Evidence on the Epidemiology of Hepatitis C Virus in Hemodialysis. Pathogens. 2021;10(9). doi: 10.3390/pathogens10091149. PMID: 34578181.
- Reddy YNV, Nunes D, Chitalia V, Gordon CE, Francis JM. Hepatitis C virus infection in kidney transplantation-changing paradigms with novel agents. Hemodial Int. 2018;22 Suppl 1(Suppl 1):S53-s60. doi: 10.1111/hdi.12659. PMID: 29694721.
- Khattab MA, Zakaria Y, Sadek E, Abd El Fatah AS, Fouad M, Khattab M, Moness HM, Adel NM, Ahmed E. Detection of hepatitis C virus (HCV) RNA in the peripheral blood mononuclear cells of HCV-infected patients following sustained virologic response. Clin Exp Med. 2023;23(1):131-40. doi: 10.1007/s10238-022-00791-7. PMID: 35066710.
- Saad ZM, Ghany WAE, Khalifa R, Higazi A, Al-Shazly M, Said M, Keryakos H. Occult HCV infection in liver transplanted patients: frequency and consequences. Clin Exp Hepatol. 2022;8(2):125-31. doi: 10.5114/ceh.2022.115116. PMID: 36092754.
- Fernández-Caso B, Fernández-Caballero J, Chueca N, Rojo E, de Salazar A, García Buey L, Cardeñoso L, García F. Infection with multiple hepatitis C virus genotypes detected using commercial tests should be confirmed using next generation sequencing. Sci Rep. 2019;9(1):9264. doi: 10.1038/s41598-019-42605-z. PMID: 31239457.
- Kim KA, Lee S, Park HJ, Jang ES, Lee YJ, Cho SB, Kim YS, Kim IH, Lee BS, Chung WJ, Ahn SH, Kim S, Jeong SH. Next-generation sequencing analysis of hepatitis C virus resistance-associated substitutions in direct-acting antiviral failure in South Korea. Clin Mol Hepatol. 2023;29(2):496-509. doi: 10.3350/cmh.2022.0345. PMID: 36880209.
- Huggett JF. The Digital MIQE Guidelines Update: Minimum Information for Publication of Quantitative Digital PCR Experiments for 2020. Clin Chem. 2020;66(8):1012-29. doi: 10.1093/clinchem/hvaa125. PMID: 32746458.
- Kojabad AA, Farzanehpour M, Galeh HEG, Dorostkar R, Jafarpour A, Bolandian M, Nodooshan MM. Droplet digital PCR of viral DNA/RNA, current progress, challenges, and future perspectives. J Med Virol. 2021;93(7):4182-97. doi: 10.1002/jmv.26846. PMID: 33538349.
Content Alerts
SignUp to our
Content alerts.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.