Biology Group. 2024 August 12;5(8):960-968. doi: 10.37871/jbres1974.
The Impact of Chloroform on Viability of Temperate Bacteriophage Lambda and Survival of its Host, Escherichia coli
Amita Pandey* and Tarannum Hashmi
- Bacteriophage lambda
- Lysogen
- Escherichia coli
- Polymerase chain reaction
- Chloroform
Abstract
Background: This study investigates the effect of chloroform, an organic compound with anti-bacterial properties, produced both naturally and due to anthropogenic activities on the viability of temperate bacteriophage lambda (phage lambda) and its host, Escherichia coli (E. coli). The study was initiated while repurposing phage lambda for testing efficacy and efficiency of disinfectants.
Results: Plaque assays and Polymerase Chain Reaction (PCR) using primers specific to the Lambda Tail Protein (LTP) demonstrated a decrease in plaque-forming units and the log copy number of phage lambda following chloroform treatment of phage lysates. Interestingly, PCR with 16 S rRNA primers detected the presence of bacteria in both chloroform-treated and untreated lysates, suggesting that E. coli lysogens, small enough to pass through 0.2 μm filters, which is commonly used for sterilization, were present in the phage lysates. The presence of lysogens in the phage lysates was further supported by the absence of a 16 S rRNA amplicon in lysates prepared without phage lambda infection and the presence of an amplicon in DNase-treated lysates. Additionally, PCR and microscopy revealed that chloroform treatment of phage lysates increased the lysogen copy number, decreased lysogen size, and altered lysogens phage-producing ability.
Conclusion: In summary, while chloroform had a slight effect on the viability of phage lambda, most of the phage remained resistant to chloroform because they lack a lipid coat. Phage lambda transitioned to a lysogenic life cycle under sub-optimal growth conditions, leading to the formation of E. coli lysogens. These lysogens were more resistant to stresses like chloroform, ensuring the survival of both the phage and the host during adverse conditions
Introduction
The temperate phage lambda was discovered in a laboratory culture of E. coli K12 strain, exhibiting lysogenic and lytic life cycles [1,2]. During the lysogenic life cycle phage lambda DNA integrates into the host genome and is called a “prophage”, and the host is called a “lysogen”. The prophage is induced and enters lytic life cycle when the lysogen encounters stressful conditions, producing mature phages and releasing them into the environment by bacterial cell lysis.
The lysogenic life cycle has been shown to be favored and regulated by CI, CII, and CIII proteins at low temperature, starvation, and high multiplicity of infection [3]. Whereas, Cro protein has been shown to regulate the lytic cycle [4]. Interestingly, CI repressor was the first protein discovered to be cleaved by recA-lexA-mediated process, involved in stress response in E. coli [5,6].
Chloroform is naturally produced and degraded by microbes [7,8]. Moreover, chloroform is a regulated Disinfection-By-Product (DBP), used in certain medical procedures, and shown to be formed during the heating and chlorination of drinking water containing traces of organic compounds [9-12].
This study was initiated to find the root cause of the anomalous results obtained during testing disinfectant efficacy against phage lambda lysates using plaque assay. It was observed that the disinfectant treated phage lysates showed higher Plaque Forming Units (PFU) compared to untreated phage lysates. It was reasoned that the increase in the PFU could be due to the presence of lysogen in the phage lysates, which were being induced by the disinfectant, which is consistent with studies showing induction of lysogens by various extrinsic and intrinsic stimuli. To test the hypothesis regarding the presence of lysogen in the phage lambda lysates, PCR assay was performed, which showed presence of E. coli lysogens in both chloroform treated and untreated lysates. Additionally, chloroform-treatment of phage lambda lysates enhanced lysogen log copy number, reduced lysogen size, and altered its phage producing ability. Furthermore, phage lambda was found to be more resistant to chloroform treatment.
Materials and Methods
Host bacterial strain
E. coli strain was used as the host bacterium for phage lambda lysate preparation and testing of disinfectants. To prepare a permissible host a single bacterial colony was inoculated into Luria Broth (0.2% Maltose and 10 mM MgSO4) and allowed to grow overnight at 37°C.
Bacteriophage
Phage lambda was a gift from Prof. GK Pandey (University of Delhi South Campus). Phage lysates were prepared by modifications of previously published methods [13,14]. For preparing phage lambda lysates, overnight culture of E. coli (0.2 mL) was infected with 0.1 mL of 1:100 dilution of phage lambda lysate with a titer of ~109 PFU. The infected bacterial cells were mixed with 3 mL of overlay agar (Luria Broth with 0.6% agar and 10 mM MgSO4) and poured onto plates containing Luria Broth A gar with 10 mM MgSO4. On the subsequent day 6 mL of SM buffer (100 mM NaCl, 8 mM MgSO4, 50 mM Tris-Cl pH 7.5, 0.01% w/v Gelatin) was added to each confluent plate and incubated overnight at 4°C. Subsequently, plates were rotated at 25°C for 1 hour at 50 rotations per minute, followed by pooling and centrifuging for 10 minutes at 10,000 revolutions per minute (rpm) at 25°C. The supernatant was transferred to fresh tubes and processed following four different methods:
Method 1 / M1 lysates: The centrifuged lysate was filter sterilized by passing through a 0.2 µm syringe filter, followed by the addition of 0.1 volume (volume/volume) of chloroform and rotation at room temperature (25°C) for 10 minutes. Subsequently, the lysate was centrifuged for 10 minutes at 10,000 rpm at 25°C. Dimethyl Sulfoxide (DMSO) was added to the supernatant to a final concentration of 7% and the lysates were stored at -80°C.
Method 2/ M2 lysates: After centrifugation, the lysate was filtered through a 0.2 µm syringe filter followed by the addition of 7% DMSO and stored at -80°C.
Method 3/ M3 lysates: To the centrifuged lysate added 0.1 volume (volume/ volume) of chloroform and rotated at room temperature for 10 minutes followed by centrifugation for 10 minutes at 10,000 rpm at 25°C. The supernatant was transferred to conical tubes and filter sterilized using 0.2 µm syringe filters. To the filtered lysate added DMSO to a final concentration of 7% and stored at -80°C.
Method 4/ M4 lysates: After centrifugation, the lysate was filtered through a 0.2 µm syringe filter and stored at -80°C.
Phage titer determination with plaque assay
The titer of phage lysates was determined by performing plaque assay with minor modifications of the published protocol [14]. Briefly, a dilution series of phage lysate (10-2, 10-4, 10-6, 10-8, 10-10) was prepared in phage dilution buffer (10 mM Tris.Cl, pH 7.5, and 10 mM MgSO4) and mixed with 0.2 mL E. coli culture in a tube containing overlay agar maintained at 45°C to 50°C and poured onto Luria Broth Agar (with 10 mM MgSO4) medium containing plates. The plates were incubated overnight at 37°C. On subsequent day, the PFUs were recorded for each dilution, and the Log10 PFU/ mL was determined.
Log copy number determination using PCR assay
For copy number determination standard curves were generated for E. coli and phage lambda DNA using 16 S rRNA and Lambda Tail Protein (LTP) primers (Supplementary table 1). Kit purified E. coli (0.8 nanogram (ng), 8 ng, and 80 ng) and phage lambda (0.005 ng, 0.05 ng, 0.5 ng, 5 ng) DNA were used as standard solutions for PCR amplification. The reaction conditions and cycling parameters for PCR amplification are provided in supplementary table 2.
For determining the copy number of phage lambda and E. coli present in the lysates, 0.006 mL of phage lysates (M1, M2, and M3) were PCR amplified with 16 S rRNA and LTP primers. For stability kinetics phage lambda lysates (M2 and M3) were diluted in phage dilution buffer (1:10) and incubated at 20°C for three days, subsequently PCR was performed with 0.006 mL of phage lysate using 16 S rRNA and LTP primers.
The PCR products of lysates and standard solutions were size separated on a 2% agarose gel slab followed by imaging and densitometry using Image J software [15]. The Percent Peak Area (PPA) obtained by densitometry was applied to calculate DNA content in nanograms (ng), which was subsequently used for calculating copy number using the following equation: [DNA (nanogram)*6.023*1023]/ [DNA (size in base pair)*109*660].
DNase treatment
Phage lysates (M2 and M3) were diluted in phage dilution buffer (1:10) and treated with 2 Units of TURBO DNase (Invitrogen) at 37°C for 30 minutes followed by heat inactivation for 20 minutes at 75°C. Subsequently 0.006 mL was used for PCR amplification.
Gram staining
For Gram-staining [16], a 0.020 mL overnight culture of E. coli and 1 mL phage lysate (M1, M2, and M3) were centrifuged for 5 minutes at 10,000 rpm at 10°C to pellet down the cells. The pellet was thereafter washed in phosphate-buffered saline (0.137M NaCl, 0.027M KCl, 0.01M Na2HPO4, 0.018M KH2PO4, pH 7.4), suspended in 0.020 mL of phosphate buffer, and spread onto glass slides as a uniform smear. The glass slides were air dried for 30 minutes. Subsequently, the dried slides were fixed by flooding with absolute methanol (2 min) and rinsing in distilled water. Thereafter, the fixed slides were stained in accordance to the manufacturer’s instructions. The slides were thereafter examined using bright-field microscopy using Olympus URFL fluorescent microscope and imaged using ProgRes® Capture Pro 2.7.7 software. Image J software was used for analyzing particle number and size. For particle number and size analysis the lower limit was set at 0 and 0.01, respectively and upper limit was set at infinity for both the parameters.
Statistical Analysis
All the plots were prepared in ggplot2 and the statistical analysis was performed using Student’s t-test [17,18]. The result was considered to be statistically significant when P values were less than .05 and highly significant when P values were less than .001.
Results and Discussion
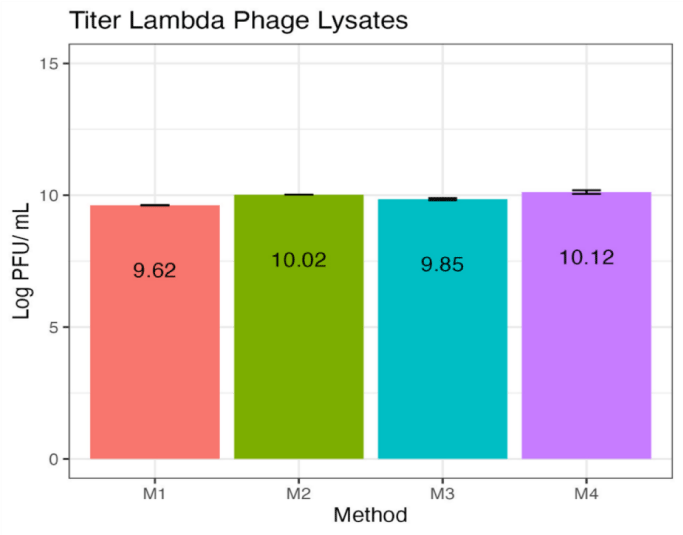
The effect of chloroform on the viability of phage lambda was initially evaluated using plaque assay (Figure 1). It was observed that phage lysates M2 and M4 prepared with the chloroform-free method exhibited higher average titers of 10.02 and 10.12 PFU/ mL, respectively. Whereas, M1 and M3 lysates prepared with methods involving chloroform-treatment showed significantly lower average titers of 9.62 and 9.85 PFU/ mL, respectively. A marginal decrease in the titer can be attributed to the lack of a lipid coat in phage lambda, which makes it more resistant to solvents like chloroform. However chloroform can still inflict some degree of protein denaturation or disruption of the capsid structure, which accounts for the observed 1 log reduction in the phage titer. These results corroborate with previous findings showing that chloroform has a destabilizing effect on phages [19]. No significant difference in the phage titers for M2 (10.02 PFU/mL) and M4 (10.12 PFU/mL) lysates was observed, which differed in addition of DMSO (Figure 1). DMSO is typically added to minimize intracellular ice formation and osmotic stress endured by cells during the freezing process that occurs upon storage at ultra-low temperatures (-80°C). These results are consistent with research showing that the cryoprotective effect of DMSO is due to the formation of water pores in the membrane, allowing the cryoprotectant to penetrate the cell and promote vitrification-a process absent in phages due to lack of membrane [20].
Interestingly, a significant difference was observed in the phage titers of chloroform-treated M1 (9.62 PFU/ mL) and M3 (9.85 PFU/ mL) lysates, which differed in the sequel of the filtration step (Figure 1). The M1 lysates were filtered before chloroform-treatment, whereas the M3 lysates were filtered both before and after chloroform-treatment. Chloroform is an antibacterial compound, which causes cell lyses and release phages. However, it can also influence properties of bacterial cell membrane, which can effect phage infection and replication, indicating that chloroform-treatment created a heterogenous population of E. coli lysogen. Inclusion of a second filtration step after chloroform-treatment preferentially filtered out some lysogens based on their morphology from the M3 lysates, while these were retained in the M1 lysates.
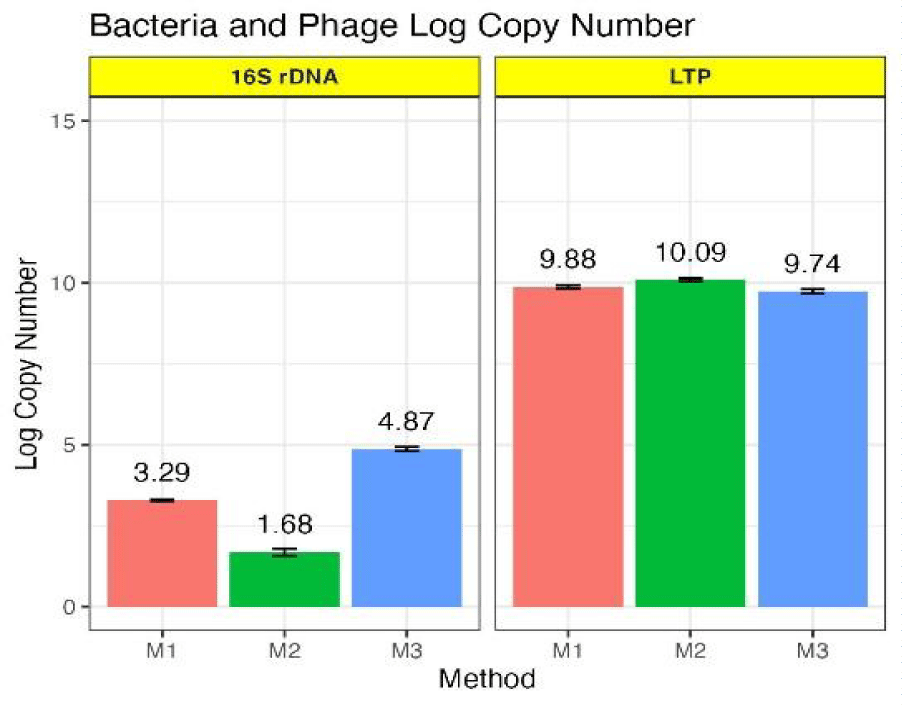
It is possible to detect presence of lysogen in lysates using plaque assay by evaluating plaque morphology. However, due to its higher sensitivity and convenience PCR assay was employed to further evaluate the composition of the phage lysates (M1, M2, and M3) by determining the log copy number of E. coli lysogen and phage lambda using 16 S rRNA and LTP primers, respectively (Figure 2). The log copy number was determined from the Percent Peak Area (PPA) of PCR bands in the gel and DNA content in nanograms calculated from the PPA as described in material and methods (Supplementary figures 1 (A-B)). The rationale behind using PPA (is positively related to the PCR band intensity in the gel) for copy number determination was that the PPA directly corresponds to the genome copy number of E. coli and phage lambda and the target gene copy number of 16 S rRNA and LTP genes present in the phage lysates. However, a corroboration was not observed between the PPA of the gel bands and the calculated DNA content for E. coli (Supplementary figure 2C). It was observed that even though the PPA was higher for phage DNA (13.47%, 14.35%, and 12.61%) and lower for E. coli DNA (4.4%, 2.23%, and 6.71%) (Supplementary figure 1B), the calculated DNA content was lower for phage lambda DNA (3.6 ng, 5 ng, 2,25 ng) and high for bacteria (21.6 ng, 3.6 ng, and 80 ng) (Supplementary figure 2C). This discord was attributed to the higher concentration of the standard solutions used for E. coli due to its larger genome size (4.6 million base pairs) compared to phage lambda for regression analysis (Supplementary figures 2 (A,B)). This resulted in an almost similar log copy number of E. coli lysogen and phage lambda present in the lysates, which was not consistent with intensity of PCR band in the gel (Supplementary figures (1A,2D)). Since the concentration of E. coli standard solutions could not be changed, therefore the true log copy number of E. coli lysogens was determined based on the ratio of the PPA for bacteria and phage DNA. The ratio of bacteria to phage PPA was calculated as 1/6 (M2), 1/3 (M1), and 1/2 (M3) and applied to calculate the true copy number. Thereafter, a corroboration was observed between the intensity of the gel bands, their PPA values, and the log copy number (Supplementary figures 1 (A,B)), and (Figure 2). Interestingly, lysogen were detected in both chloroform-treated and untreated lysates using a PCR assay which is consistent with research showing that bacteria form lysogen in the presence of temperate phages when exposed to adverse conditions [3]. Moreover, the chloroform-treated lysates (M1 and M3) showed significantly higher copy number of lysogen compared to untreated lysates, which could be a contributing factor to the observed differences in the PFU/ mL (Figure 1). Detection of lysogen in chloroform-treated lysates was an unexpected result given the known antibacterial property of chloroform [19]. Contrarily, M1 and M3 lysates showed significantly higher lysogen copy number compared to M2 lysates. These observations indicate that the lysogen might be more resistant to chloroform-treatment and chloroform might be inducing stress responses enabling lysogen to survive and proliferate. These conclusions are supported by research showing that prophage enables bacteria to encounter environmental stresses [21,22].
Additionally, the log copy number values of phage lambda determined with the PCR method were similar to the PFU/ mL values obtained from plaque assay (Figures (1,2)). These findings corroborate with studies showing application of real-time PCR for quantitation of phages [23]. This study shows that a PCR assay can reliably quantitate phages and bacteria in the lysates.
To further support presence of E. coli lysogens in the lysates, phage lysate was prepared using method 3 in absence of phage lambda infection. No bacterial DNA was detected in this lysate using 16 S rRNA primers, indicating that the bacterial DNA found in the M1, M2, and M3 lysates originated from the lysogen. Presence of lysogens in the lysates was further confirmed by performing PCR assay with DNase-treated M2 and M3 lysates (Supplementary figure 3). The rationale being that PCR might be amplifying DNA present in the solution, released upon cell lysis caused by various stresses including chloroform treatment and adverse growth conditions. Evaluation of the percent peak area of DNase-treated M3T (1.63%) and M2T (2.03%) lysates showed the presence of 16 S rRNA amplicon indicating the presence of intact E. coli lysogens. Since DNase-treatment included incubations at 37°C and 70°C, therefore the data was not used for further analysis as such temperatures can induce the lysogens and reduce phage viability, resulting in misinterpretation of results.
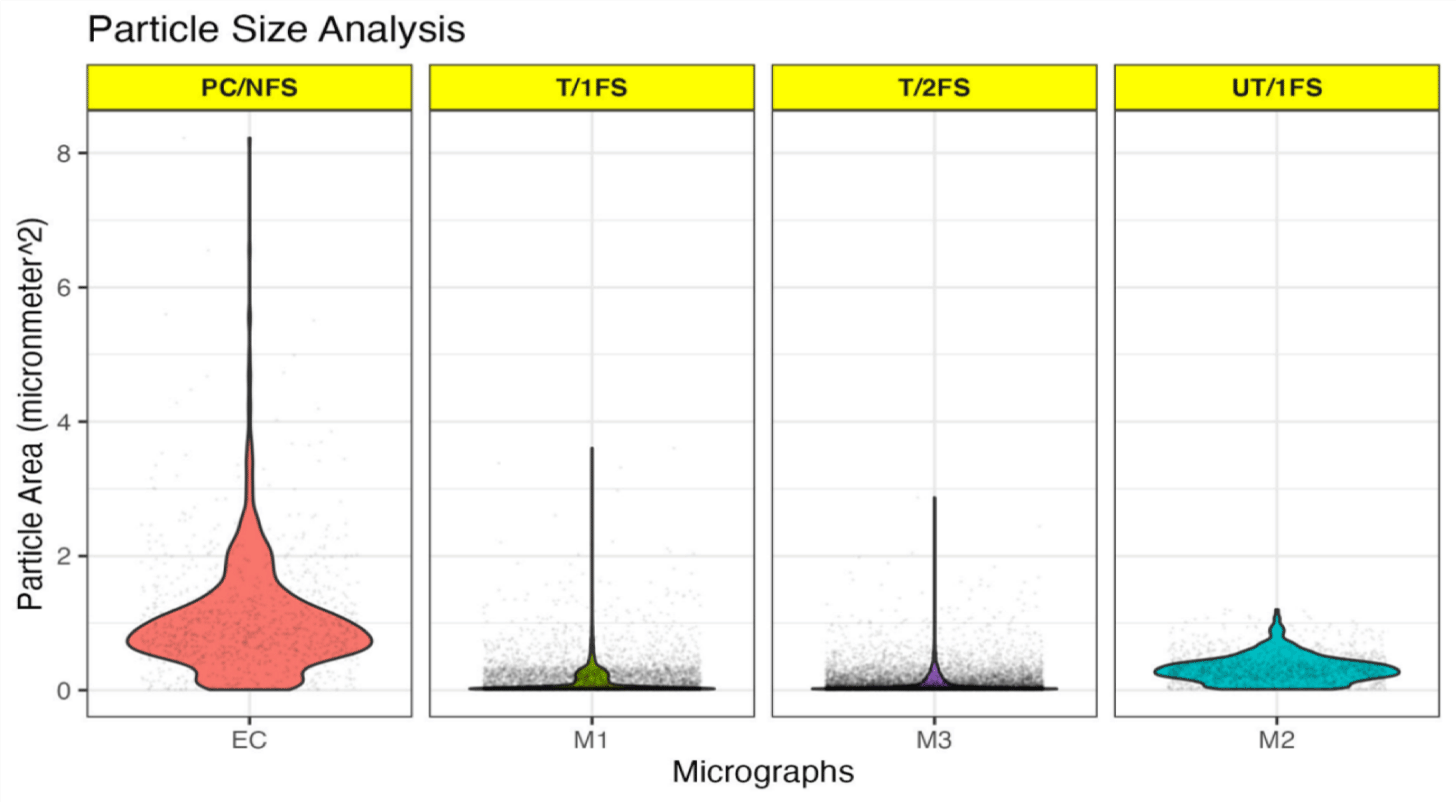
Since the lysogen were able to pass through 0.2 μm filters, generally used for filter sterilization of lysates to filter-out bacteria, gram-staining was performed to estimate the size and number of lysogen present in the lysates (M1, M2, and M3) (Supplementary figure 4A). Rod-shaped E. coli cells were visible in the positive control, whereas the lysates showed the appearance of smaller spherical pink bodies/ particles. A significantly fewer spherical bodies were observed for the untreated M2 lysate (526 ± 185) compared to chloroform-treated M1 (2304 ± 1135) and M3 (3670 ± 2378) lysates corroborating with the log copy number values for E. coli lysogen, indicating that the spherical bodies were not an artifact of gram-staining but due to inherent differences in the composition of the lysates due to presence of lysogen (Supplementary figure 4B). No significant difference in the lysogen number was observed between the chloroform-treated M1 and M3 lysates. For size determination the average area (μm2) was analyzed and found to be highest for M2 (0.302 ± 0.08) followed by M1 (0.148 ± 0.04) and M3 (0.111 ± 0.02) lysates (Figure 3). The lysogen present in chloroform-treated (M1 and M3) lysates were significantly smaller than the untreated (M2) lysates. Although, no significant size difference was observed between M1 and M3 lysates, careful visual examination of the plots did reveal presence of heterogeneous population of lysogen. The lysogen in the M1 lysates were visibly bigger than the lysogen present in the M3 lysates probably due to inclusion of a second filtration step after chloroform treatment while preparing M3 lysates. The average area of E. coli cells (no filtration step involved) was 1.04, although smaller and larger cells were also detected, which is consistent with studies showing that bacteria undergo morphological changes upon encountering stress [24]. In conclusion, the increase in number and decrease in size of lysogens upon chloroform-treatment indicated that chloroform might be exerting additional stress, which might induce stress responses leading to the observed morphological due to ongoing physiological changes.
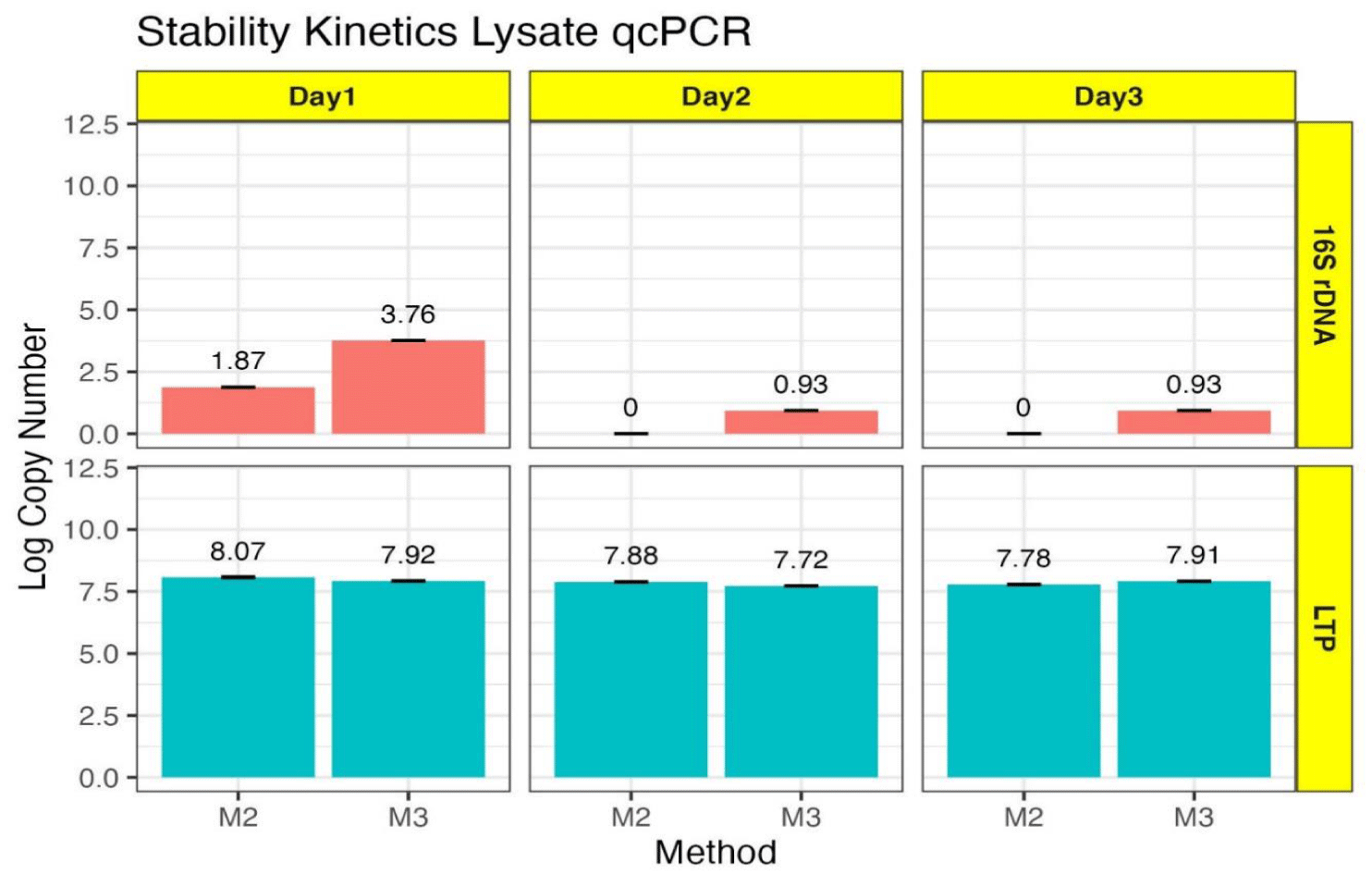
The effect of chloroform on the physiology of the lysogens was further analyzed by performing a three-day stability kinetics of the lysates (M2 and M3) using PCR assay (Figure 4). Interestingly, the copy number of lysogen showed significant reduction from Day1 to Day2 for both M2 (1.87 to 0) and M3 (3.76 to 0.93). Thereafter, no reduction in copy number was observed for M3 lysates. Since a 16 S rRNA amplicon was not detectable in M2 lysates on Day2 and Day3 under these PCR conditions it could not be analyzed any further. The initial decrease in the copy number of the lysogen might be due to stress responses induced by the change in environmental conditions i.e. shift from ultra-low temperature to room temperature. Such stress can activate bacterial defenses or lead to an initial destabilization of the prophage within the host genome. Over time, the bacterium may adapt to the new temperature conditions, leading to the stabilization of the prophage DNA replication mechanisms, which is evident as no significant reduction in lysogen copy number was observed for M3 lysates from Day2 to Day3. As the cells recover, the prophage might also start replicating more efficiently. The phage lambda copy number exhibited a significant decrease from Day1 (7.92) to Day2 (7.72) followed by increase on Day3 (7.91) for M3 lysates. Whereas, a significant reduction was observed for M2 lysates: Day1 (8.07), Day2 (7.88), and Day3 (7.78). These results indicate an inherent difference in phage producing ability between chlroform treated and untreated lysates. The observed fluctuation in phage lambda copy number also indicates that the host bacterium and the prophage are undergoing a period of adjustment to the new environmental conditions. Moreover, chloroform-treatment can cause additional stress to the bacterial cells, potentially leading to DNA damage or other forms of cellular stress that can affect prophage stability and replication accounting for the difference in the observed stability kinetics for M2 and M3 lysates. These conclusions are supported by research showing that prophage undergo both induced and spontaneous induction regulated by intrinsic and extrinsic factors and is beneficial for the host bacterium to survive through adverse conditions [25].
Conclusion
This study shows that the phage lambda is more resistant to chloroform treatment due to absence of a lipid coat. Although, chloroform might inflict minimal damage to its capsid coat leading to some loss in viability. Moreover, owing to its temperate nature phage lambda underwent a lysogenic life cycle resulting in formation of E. coli lysogen, which could easily pass through the traditionally used 0.2 μm filters employed for filter sterilization purpose. This underscores the need for better filter sterilization techniques, since presence of lysogen might interfere downstream applications such as disinfectant testing, isolating high quality phage lambda DNA, etc. Contrary to its role as an antibacterial compound, chloroform treatment enhanced lysogen copy number, reduced lysogen size, and altered their stability and phage producing ability, indicating that lysogen might be more resistant to stresses enabling E. coli to overcome adverse conditions. These findings are supported by research showing that lysogen can exhibit increased resistance to certain types of stress compared to non-lysogenic bacteria attributed to several factor including superinfection immunity, resistance genes, stabilization of host genome, and toxin-antitoxin systems [26].
Acknowledgment
The authors extend sincere thanks to the management of Shriram Institute for Industrial Research for providing infrastructure facilities including equipment, reagents, and chemicals, which were required for completion of this work. A special thanks is extended to Dr. Mukul Das (Director) for supporting the work and taking time from his busy schedule to read and approve the manuscript (SRI-MS# 20240102-01).
Funding
This work received no specific grant from any funding agency.
Conflicts of interest/Competing interests
None reported Ethics approval: Not required.
Consent to participate
All others have contributed to the manuscript preparation.
Consent for publication
All authors have read and approved the manuscript.
Availability of data and material
Data is available on request.
Author contribution
Amita conceived the study idea and design, performed quantitative conventional PCR, gram staining, microscopy, plaque assays, collected data, performed analysis, and prepared the manuscript. Tarannum performed plaque assays, gram staining, and PCR amplification.
References
- LEDERBERG J. Streptomycin resistance; a genetically recessive mutation. J Bacteriol. 1951 May;61(5):549-50. doi: 10.1128/jb.61.5.549-550.1951. PMID: 14832197; PMCID: PMC386043.
- Hershey AD. The bacteriophage lambda. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory; 1971.
- Oppenheim AB, Kobiler O, Stavans J, Court DL, Adhya S. Switches in bacteriophage lambda development. Annu Rev Genet. 2005;39:409-29. doi: 10.1146/annurev.genet.39.073003.113656. PMID: 16285866.
- Folkmanis A, Takeda Y, Simuth J, Gussin G, Echols H. Purification and properties of a DNA-binding protein with characteristics expected for the Cro protein of bacteriophage lambda, a repressor essential for lytic growth. Proc Natl Acad Sci U S A. 1976 Jul;73(7):2249-53. doi: 10.1073/pnas.73.7.2249. PMID: 1065873; PMCID: PMC430516.
- Little JW. Autodigestion of lexA and phage lambda repressors. Proc Natl Acad Sci U S A. 1984 Mar;81(5):1375-9. doi: 10.1073/pnas.81.5.1375. PMID: 6231641; PMCID: PMC344836.
- Walker GC. The SOS Response of Escherichia coli. In: Neidhardt FC, Curtiss R III, Ingraham JL, Lin EC, Low KB, Magasanik B, Reznikoff WS, Riley M, Schaechter M, Umbarger HE, editors. Escherichia coli and Salmonella: Cellular and molecular biology. Washington, D.C.: American Society for Microbiology; 1996. p.1400-1416.
- Cappelletti M, Frascari D, Zannoni D, Fedi S. Microbial degradation of chloroform. Appl Microbiol Biotechnol. 2012 Dec;96(6):1395-409. doi: 10.1007/s00253-012-4494-1. Epub 2012 Oct 24. PMID: 23093177.
- Weigold P, Ruecker A, Jochmann M, Osorio Barajas XL, Lege S, Zwiener C, Kappler A, Behrens S. Formation of chloroform and tetrachloroethene by Sinorhizobium meliloti strain 1021. Lett Appl Microbiol. 2015 Oct;61(4):346-53. doi: 10.1111/lam.12462. Epub 2015 Aug 4. PMID: 26119060.
- Wawersik J. Die Geschichte der Chloroformnarkose [History of chloroform anesthesia]. Anaesthesiol Reanim. 1997;22(6):144-52. German. PMID: 9487785.
- Good ML, McCammon A. An removal of gutta-percha and root canal sealer: a literature review and an audit comparing current practice in dental schools. Dent Update. 2012 Dec;39(10):703-8. doi: 10.12968/denu.2012.39.10.703. PMID: 23367635.
- Forczek ST, Pavlík M, Holík J, Rederer L, Ferenčík M. The natural chlorine cycle - Formation of the carcinogenic and greenhouse gas compound chloroform in drinking water reservoirs. Chemosphere. 2016 Aug;157:190-9. doi: 10.1016/j.chemosphere.2016.05.017. Epub 2016 May 24. PMID: 27231877.
- Li XF, Mitch WA. Drinking Water Disinfection Byproducts (DBPs) and Human Health Effects: Multidisciplinary Challenges and Opportunities. Environ Sci Technol. 2018 Feb 20;52(4):1681-1689. doi: 10.1021/acs.est.7b05440. Epub 2018 Jan 21. PMID: 29283253.
- Patterson TA, Dean M. Preparation of high titer lambda phage lysates. Nucleic Acids Res. 1987 Aug 11;15(15):6298. doi: 10.1093/nar/15.15.6298. PMID: 2957644; PMCID: PMC306089.
- Sambrook J, Fritsch ER, Maniatis T. Molecular Cloning: A Laboratory Manual. 2nd ed. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989.
- Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012 Jul;9(7):671-5. doi: 10.1038/nmeth.2089. PMID: 22930834; PMCID: PMC5554542.
- Gram HC. Über die isolierte Färbung der Schizomyceten in Schnitt- und Trockenpräparaten. Fortschritte der Medizin (in German). 1884;2:185-189.
- Student. The probable error of a mean. Biometrika. 1908;1-25.
- Wickham H. ggplot2: Elegant Graphics for Data Analysis. 2nd ed. Springer-Verlag New York; 2016.
- Taylor J, Bettelheim KA. The action of chloroform-killed suspensions of enteropathogenic Escherichia coli on ligated rabbit-gut segments. J Gen Microbiol. 1966 Feb;42(2):309-13. doi: 10.1099/00221287-42-2-309. PMID: 5330337.
- Whaley D, Damyar K, Witek RP, Mendoza A, Alexander M, Lakey JR. Cryopreservation: An Overview of Principles and Cell-Specific Considerations. Cell Transplant. 2021 Jan-Dec;30:963689721999617. doi: 10.1177/0963689721999617. PMID: 33757335; PMCID: PMC7995302.
- Brüssow H, Canchaya C, Hardt WD. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol Mol Biol Rev. 2004 Sep;68(3):560-602, table of contents. doi: 10.1128/MMBR.68.3.560-602.2004. PMID: 15353570; PMCID: PMC515249.
- Fortier LC, Sekulovic O. Importance of prophages to evolution and virulence of bacterial pathogens. Virulence. 2013 Jul 1;4(5):354-65. doi: 10.4161/viru.24498. Epub 2013 Apr 23. PMID: 23611873; PMCID: PMC3714127.
- Edelman DC, Barletta J. Real-time PCR provides improved detection and titer determination of bacteriophage. Biotechniques. 2003 Aug;35(2):368-75. doi: 10.2144/03352rr02. PMID: 12951778.
- van Teeseling MCF, de Pedro MA, Cava F. Determinants of Bacterial Morphology: From Fundamentals to Possibilities for Antimicrobial Targeting. Front Microbiol. 2017 Jul 10;8:1264. doi: 10.3389/fmicb.2017.01264. PMID: 28740487; PMCID: PMC5502672.
- Nanda AM, Thormann K, Frunzke J. Impact of spontaneous prophage induction on the fitness of bacterial populations and host-microbe interactions. J Bacteriol. 2015 Feb;197(3):410-9. doi: 10.1128/JB.02230-14. Epub 2014 Nov 17. PMID: 25404701; PMCID: PMC4285972.
- Imamovic L, Ballesté E, Jofre J, Muniesa M. Quantification of Shiga toxin-converting bacteriophages in wastewater and in fecal samples by real-time quantitative PCR. Appl Environ Microbiol. 2010 Sep;76(17):5693-701. doi: 10.1128/AEM.00107-10. Epub 2010 Jul 9. PMID: 20622134; PMCID: PMC2935055.
Content Alerts
SignUp to our
Content alerts.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.