Medicine Group. 2024 August 05;5(8):880-895. doi: 10.37871/jbres1967.
Mechanistic Insights into the Anti-Osteoporotic Effects of Naringin via Network Pharmacology and Molecular Docking Studies
Xiaodong Li1†, Xin Qi2†, Yincang Wang2, Xinyue Yang3, Xiangyu Cui1, Dawang Wang1, Boyuan Kang1, Xiaofeng Zhang4* and Xilin Xu1*
2Heilongjiang University of Chinese Medicine, Harbin 150040, China
3The second Affiliated Hospital of Heilongjiang University of Chinese Medicine, Harbin 150040, China
4Heilongjiang Administration of Traditional Chinese Medicine, Harbin 150036, China
†These authors contributed equally to this work
- Osteoporosis
- Naringin
- Apoptosis
- Network pharmacology
- Pathway
- Target
- Molecular docking
Abstract
Background: Naringin, a natural flavonoid prevalent in citrus fruits, is known for its antioxidant, anti-inflammatory, and antitumor properties. Recent studies suggest its potential influence on bone metabolism, proposing it as an anti-osteoporotic agent. However, specific mechanisms of action remain unclear.
Objective: This study aimed to validate the anti-osteoporotic effects of naringin by combining network pharmacology analysis and molecular docking validation.
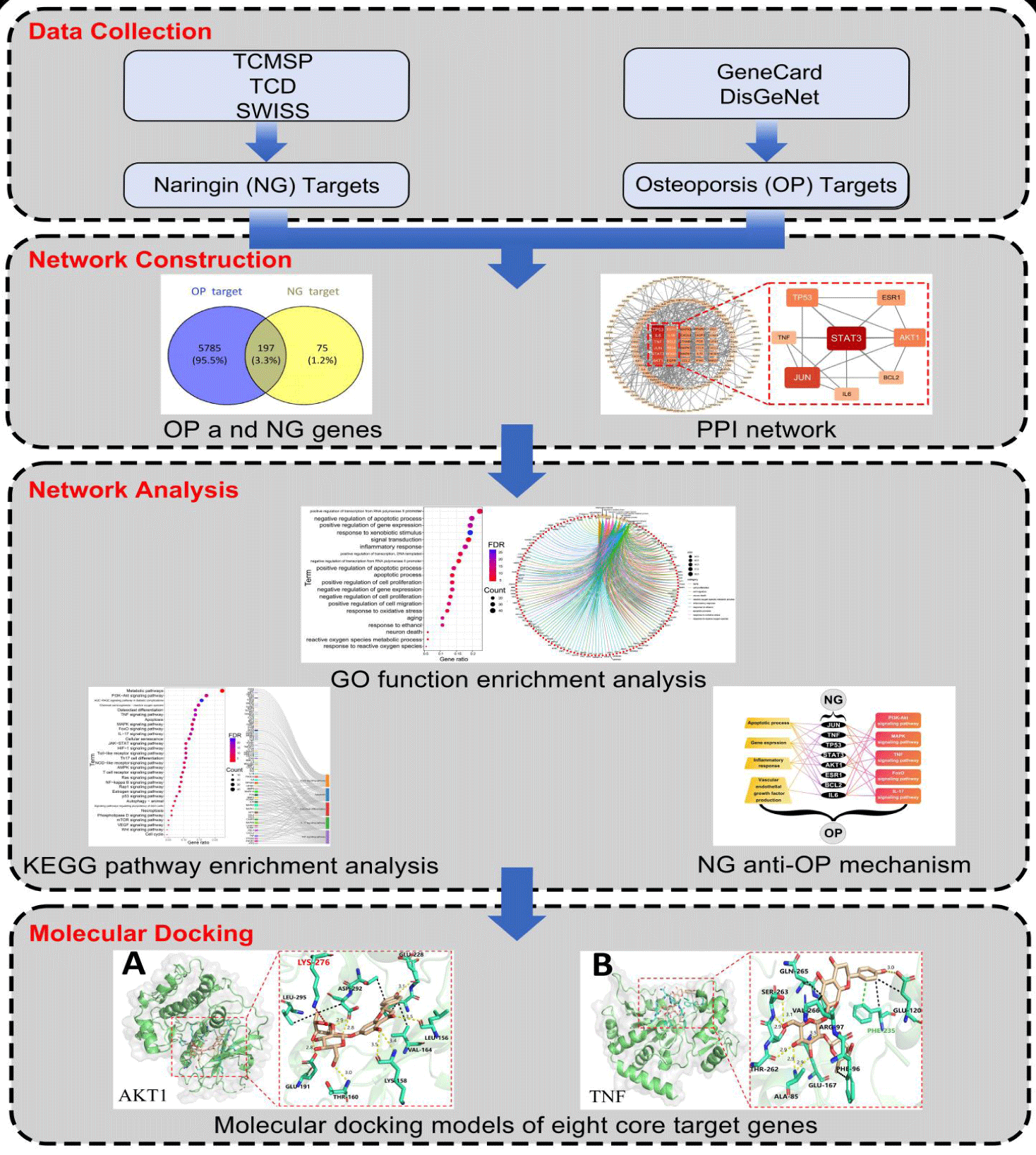
Methods: Employing network pharmacology and molecular docking, we constructed an analysis framework for "Naringin-Targets-GO and KEGG-Osteoporosis" using Cytoscape software and R software. This framework integrated various online public databases, Protein-Protein Interaction (PPI) analysis, Gene Ontology (GO) analysis, Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis, and molecular docking using AutoDock Vina software.
Results: We identified 272 target genes associated with naringin and 5,785 target genes related to osteoporosis, with 197 overlapping genes. Further analysis led to the selection of eight core targets: TP53, TNF, JUN, STAT3, AKT1, ESR1, BCL2, and IL6. GO functional enrichment analysis and KEGG pathway enrichment analysis revealed that naringin might influence osteoporosis progression by regulating biological processes, such as apoptosis, cell proliferation, and inflammatory responses, as well as signaling pathways, such as TNF, FoxO, and PI3K-Akt. Molecular docking validation confirmed the high binding affinity of naringin to the eight core targets, with binding energies below 7.2 kcal/mol.
Conclusion: The therapeutic mechanism of naringin against osteoporosis may involve the regulation of gene expression, including TP53, IL6, TNF, JUN, STAT3, AKT1, ESR1, and BCL2, and mediating signaling pathways such as TNF, FoxO, IL-17, PI3K-Akt, AMPK, and VEGF, thereby affecting multiple biological processes, such as bidirectional regulation of osteoblast proliferation and apoptosis, angiogenesis, and downregulation of inflammatory responses.
Abbreviations
NG: Naringin; OP: Osteoporosis; GO: Gene Ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes; TCM: Traditional Chinese Medicine; PPI: Protein-Protein Interaction Network; FDR: False Discovery Rate; BP: Biological Process; MF: Molecular Function; CC: Cell Composition
Introduction
Osteoporosis (OP) is a prevalent systemic skeletal disease characterized by disrupted bone homeostasis mediated by osteoclasts and osteoblasts, resulting in reduced bone mass and deterioration of bone tissue microstructure, thereby increasing bone fragility and fracture risk [1]. With global population aging intensifying, OP's prevalence is rapidly rising, posing a significant public health challenge [2]. Pharmacological treatment is the mainstay for OP; however, medications inhibiting bone resorption can lead to numerous adverse effects, limiting their long-term use [3]. Therefore, there is an urgent need for novel therapeutics for OP.
Naringin (NG), a natural flavonoid compound found in traditional Chinese medicinal ingredients such as Fructus Aurantii and Bitter Orange, exhibits various pharmacological effects, primarily in antioxidant, anti-inflammatory, and antitumor therapies [4]. Recent animal studies have demonstrated that NG modulates bone metabolism and mitigates oxidative stress via the NF-κB, PI3K/Akt, and Wnt signaling pathways [5-8], suggesting that NG may have the potential to be a novel drug for patients with OP. In Traditional Chinese Medicine (TCM), osteoporosis falls under "bone atrophy," and while TCM shows significant therapeutic effects on OP, the molecular mechanisms remain unclear [9].
Leveraging advancements in computational analysis and virtual screening, this study combined network pharmacology and molecular docking techniques to preliminarily analyze the mechanism of NG in combating osteoporosis. By utilizing public databases and analytical platforms, we constructed a "Natural compound-Gene-Target-Disease" network system to predict NG's potential core targets and pathways against OP. Furthermore, employing a 'natural compound-core target' model, we analyzed core targets to assess docking results specifics, including affinity and binding energy [10]. This study aims to provide a theoretical foundation for TCM's research in treating osteoporosis by comprehensively and deeply analyzing NG's anti-OP mechanisms through network pharmacology and molecular docking from multiple perspectives.
Data Collection and Analysis
Public databases and websites
Experimental data in this study were collected and analyzed using the databases detailed in table 1.
| Table 1: Public databases and websites. | ||
| Database Name | Website | Function |
| TCMSP Database | https://tcmsp-e.com/tcmspsearch.php | Acquire targets of NG along with their 2D and 3D structures |
| TCD Database | https://ctdbase.org/ | Obtain targets and their functions |
| PubChem Database | https://pubchem.ncbi.nlm.nih.gov/ | search for the Smile molecular formulas |
| Swiss Database | https://www.swisstargetprediction.ch/ | Gather drug targets |
| GeneCard Database | https://www.genecards. org/ | compile disease-associated targets |
| DisGeNet Database | https://www.disgenet.org/ | compile disease-associated targets |
| String Database | https://cn.string-db.org | Data analysis |
| David Database | https://david.ncifcrf.gov/ | Extract enriched information |
| Kegg Database | https://www.genome.jp/kegg/ | Obtain signaling pathway diagrams |
| Venny 2.1 | https://bioinfogp.cnb.csic.es/tools/venny/index.html | Create Venn diagrams for data visualization |
| PDB Database | https://www.rcsb.org/ | Search for and download 3D structures of the identified targets |
Construction of the naringin target database
Potential targets of NG were identified and retrieved from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) [11] and the Traditional Chinese Medicine Database (TCD) Topology [12], along with specific actions of NG targets. Furthermore, the PubChem Database [13] provides the SMILES notation for NG, which was subsequently imported into the Swiss Target Prediction Database [14] to identify additional potential NG targets. Using the STRING Database [15], protein targets corresponding to NG were converted into human genes, thereby constructing a network illustrating the relationship between NG as a single compound and its target genes.
Establishment of an osteoporosis-related disease target database and intersection targets with naringin
The keyword "Osteoporosis" was input into the GeneCards Databases [16] and DisGeNet Databases [17] to merge disease targets associated with OP and remove duplicates to obtain the final targets of OP. These targets were then analyzed using Venny 2.1 [18] to create a Venn diagram comparing the action targets of NG with OP target genes, identifying common targets as the intersection targets of Naringin-Osteoporosis (Figure 1,B).
Construction of a protein-protein interaction network for common targets and topological analysis
The common targets identified in section 1.3 were imported into the STRING database to generate a protein-protein interaction network with a confidence score of 0.4. The resulting data were saved in tsv format and visualized using Cytoscape 3.9.1 software [19], where nodes represent common targets and edges depict their interconnections. The node size and color were adjusted based on the degree value, with higher degree values corresponding to larger node sizes and darker colors [20]. A bar chart displaying the relevant targets was generated using R software [21]. For topological analysis, the CytoNCA [22] plugin within Cytoscape was employed to filter core targets based on degree values, resulting in the creation of separate diagrams for each [23].
Gene Ontology (go) functional analysis and kyoto encyclopedia of genes and genomes (kegg) pathway analysis with visualization
The common targets identified in section 1.3 underwent online analysis using the Database for Annotation, Visualization, and Integrated Discovery (David) [24], with "Official Gene Symbol" selected and the species specified as "Homo sapiens." Subsequently, the analytical data were visualized for enrichment information using R software packages such as "clusterProfiler,""ggplot2," "GOplot," and "circlize." Entries with a corrected P-value below the threshold of 0.05 were selected and ordered based on their adjusted P-values. Top-ranking entries were imported into RStudio 4.3.2 software for GO annotation and KEGG pathway enrichment analysis.
Molecular docking of core targets
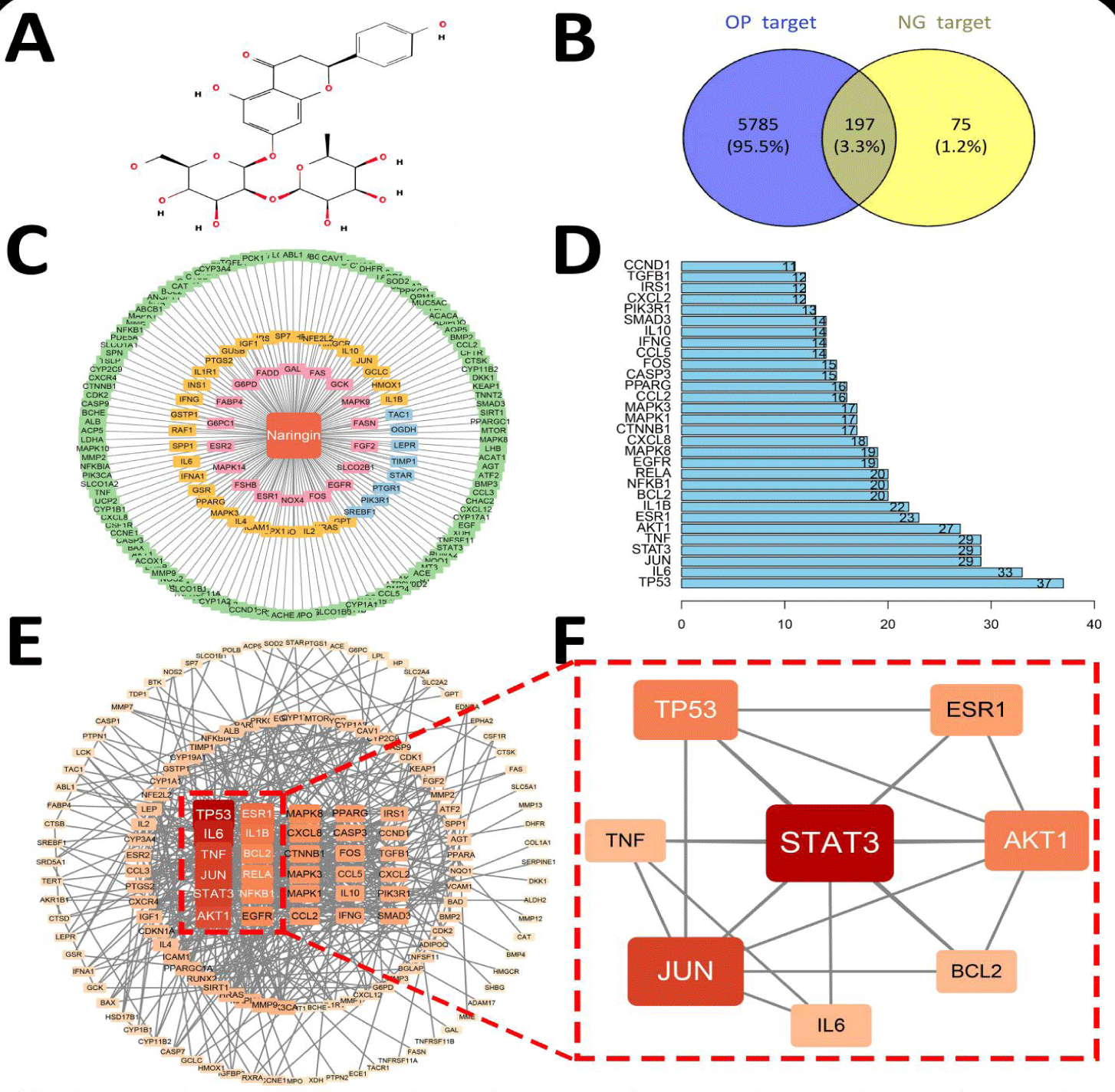
The 2D structure image of Naringin (NG) was initially retrieved from the TCMSP database (Figure 2A). Subsequently, the 3D molecular conformation of NG was downloaded in the mol2 format. Upon opening the 3D structure with AutoDockTools-1.5.7 [25], hydrogen atoms were supplemented, and charges were computed. Torsion centers and bonds were calibrated as ligands and exported in the pdbqt format. Core target proteins, chosen based on the degree value in section 1.4, had their macromolecular 3D structures acquired from the String database and the RCSB PDB database [26]; adhering to specific criteria: 1) Receptor proteins sourced from Homo sapiens, 2) Resolution generally below 3 Å, and 3) Presence of a small molecule ligand in the complex to ensure receptor protein integrity. Within the Schrödinger software [27], the macromolecule of the core target underwent removal of water molecules and solvent impurities, followed by protein separation. The Protein Preparation Wizard module facilitated preliminary processing, including residue repair and hydrogen addition, with the receptor then charged and saved in the pdbqt format. The processed NG ligand and macromolecular receptor were imported into AutoDockTools to establish the Grid Box parameters for molecular docking, ensuring appropriate enclosure of the receptor protein. Docking parameters were saved in the txt format. Using these specifications, the receptor protein was docked with the original ligand and the docking scores were compared. AutoDock Vina [28] was executed using the txt file as the working directory to initiate virtual docking. Ultimately, PyMOL-4.3 [29] software was used to the docking results of the aforementioned targets.
Results
Screening of potential targets of naringin for osteoporosis
The potential targets of NG were screened using three databases: TCMSP, CTD, and Swiss. A total of 5,179 and 270 predicted targets were collected, resulting in 272 targets being identified after removing duplicates. For OP, related gene targets were identified using DisGeNet and GeneCard Database, yielding 1,098 and 5,741 targets, respectively. After merging and removing duplicates, a total of 5,785 related genes were obtained. Finally, the intersection of NG and OP targets was visualized using Venny 2.1, identifying 197 common target genes (Figure 2B-D).
"Naringin-Action Targets" Network and Protein-Protein Interaction Network
The 197 potential targets of NG against OP were imported into Cytoscape 3.9.1 software to construct the "Naringin-Target-Action" network. This network diagram features a single NG node (highlighted in red) with the remaining nodes representing various targets. Green nodes indicate NG's inhibitory effect on corresponding gene-related proteins, pink nodes suggest NG's promotional effect, yellow nodes represent the dual action of NG on these proteins, and blue nodes indicate an interaction between NG and the interplay among multiple action targets. The shared target genes' protein-protein interaction (PPI) network was filtered using the CytoNCA plugin to select network nodes based on topological parameters such as Degree Centrality (DC), Betweenness Centrality (BC), and Closeness Centrality (CC). The median values of these parameters were obtained, and targets below the median were removed. After two rounds of analysis, the core gene network was identified, ranking TP53, IL6, TNF, JUN, STAT3, AKT1, ESR1, and BCL2 as the top eight core target genes (Table 2). The network diagram illustrating the interconnections between the core targets was constructed, featuring 8 nodes and 20 edges (Figure 2E,F). This suggests that these targets may be closely related to the mechanism of NG's anti-osteoporotic action. To further elucidate the degrees of freedom associated with the key genes, we used the R software to compute the relevant values. These were subsequently presented in a bar chart for the top 30 targets (Figure 2D).
| Table 2: Analysis Results of Osteoporosis-Naringin. | ||||
| Target Gene | Target Protein | Degree | Betweenness | Closeness |
| TP53 | Cellular tumor antigen p53 | 37 | 5018.9683 | 0.43801653 |
| IL6 | Interleukin-6 | 33 | 3782.3745 | 0.4151436 |
| TNF | Tumor necrosis factor | 31 | 1607.9734 | 0.3955224 |
| JUN | Transcription factor Jun | 29 | 3261.53 | 0.44044322 |
| STAT3 | Signal transducer and activator of transcription 3 | 29 | 2281.9084 | 0.42741936 |
| AKT1 | RAC-alpha serine/threonine-protein kinase | 27 | 1415.2236 | 0.4151436 |
| ESR1 | Estrogen receptor | 23 | 2405.9062 | 0.40561223 |
| BCL2 | Apoptosis regulator Bcl-2 | 20 | 2296.4863 | 0.39650872 |
GO functional enrichment analysis of "naringin-osteoporosis"
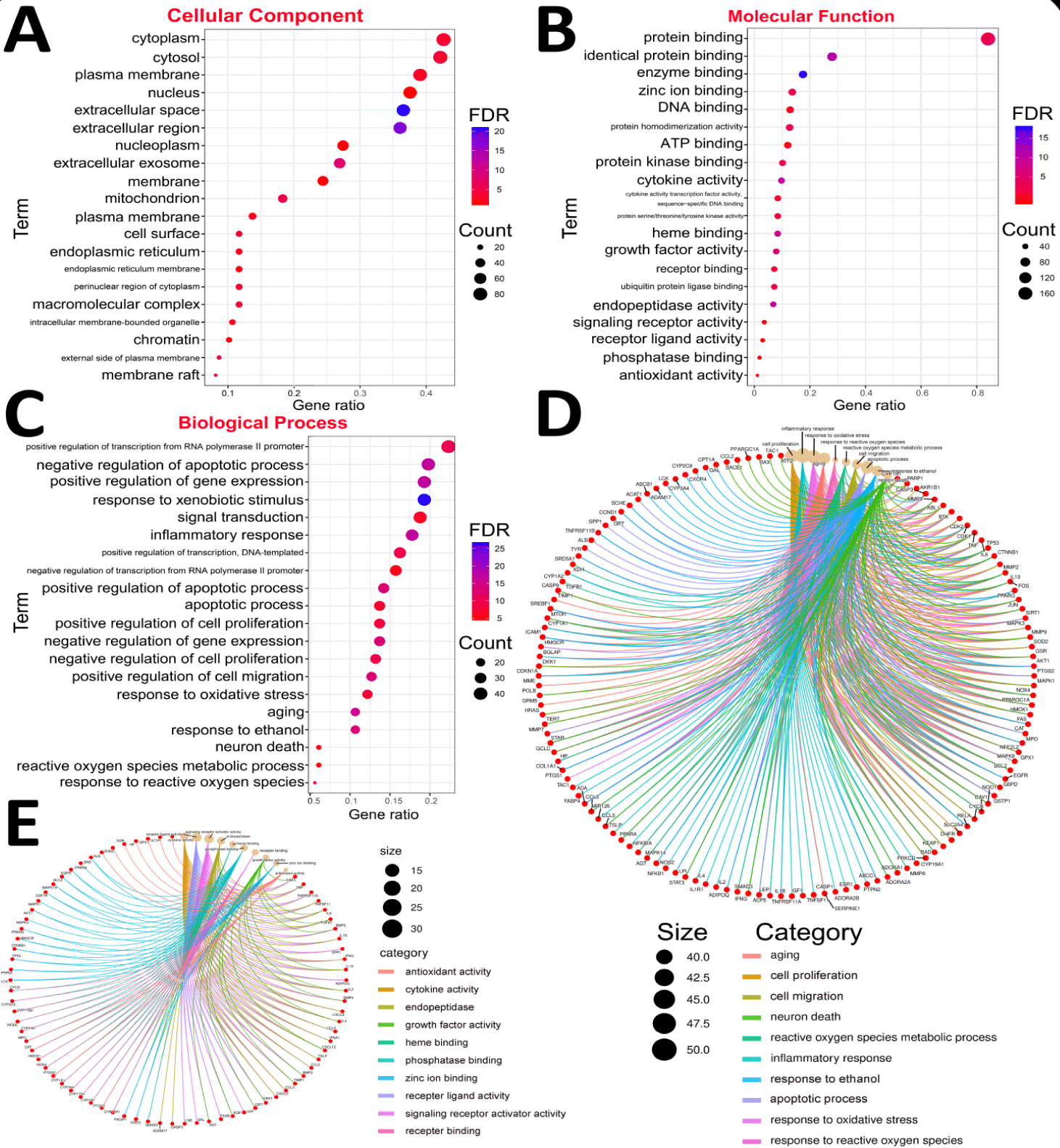
To delve deeper into the potential mechanisms of NG in treating OP, GO enrichment analysis was conducted using the David online analytical platform. The analysis yielded 1,355 enriched items, and 197 of the 197 genes were annotated. Significant enrichment was identified in the Cellular Component (CC) category, with a False Discovery Rate (FDR) value of less than 0.05, encompassing processes such as cytoplasm (GO:0005737), cytosol (GO:0005829), cell membrane (GO:0005886), nucleus (GO:0005634), and extracellular space (GO:0005615), etc, totaling 29items (Figure 3A). In the Molecular Function (MF) category, enrichment was observed in 61 items, including protein binding (GO:0005515), identical protein binding (GO:0042802), enzyme binding (GO:0019899), zinc ion binding (GO:0008270), and protein homodimerization activity (GO:0042803), etc. (Figure 3B). Within the Biological Process (BP) category, 730 items were significantly enriched, including positive regulation of transcription from the RNA polymerase II promoter (GO:0045944), negative regulation of apoptotic processes (GO:0043066), response to xenobiotic stimulus (GO:0009410), positive regulation of gene expression (GO:0010628), and signal transduction (GO:0007165), etc. (Figure 3C,D). The items were first filtered by the False Discovery Rate (FDR) values in ascending order, selecting the top 20 items in Biological Processes (BP), Cellular Components (CC), and Molecular Functions (MF). These were visualized using significant bubble charts. Building on this foundation, we further explored the most significantly expressed Gene Ontology (GO) categories within the dataset to uncover the regulatory mechanisms underlying key biological processes, molecular functions, and cellular components. Considering the biological relevance and specific characteristics of the dataset, we identified ten GO categories with distinct expression patterns. Therefore, we focused on the associated BP and MF categories and created a Chord Diagram for effective visualization (Figures 3(E,F)).
KEGG pathway enrichment analysis of "naringin-osteoporosis"
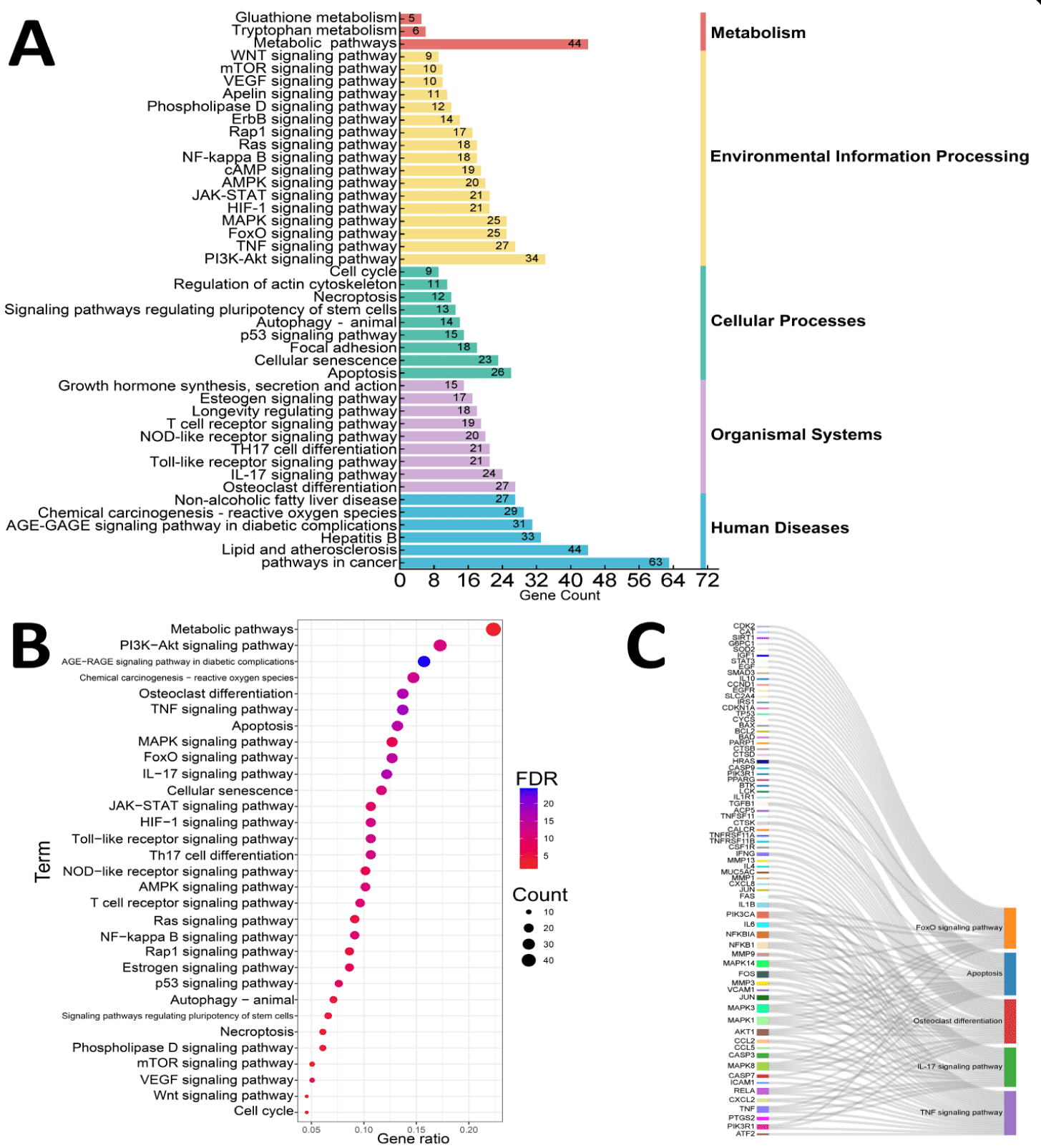
The Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of the common targets between NG and OP, using a FDR threshold of less than 0.05, identified 164 significant signaling pathways. These pathways included metabolic pathways (hsa:01100), the PI3K-AKT signaling pathway (hsa:04151), Chemical carcinogenesis-reactive oxygen species (hsa:05208), Apoptosis (hsa:04210), and osteoclast differentiation (hsa:04380), etc. To conduct an in-depth analysis and visualize the signaling pathways, we selected pathways previously reported in the literature and systematically categorized them into three hierarchical levels based on the classification criteria of KEGG official website. Seven major categories were identified at the highest level of classification. Through meticulous classification analysis of the KEGG database, the study identified the following pathways: 6 metabolic pathways, 23 signaling pathways, 12 cellular processes, 45 organismal systems, and 45 human diseases [30]. To effectively understand the complex signaling pathways within the body, we selected 45 pathways for visual analysis of KEGG pathway classification (Figure 4A). Relevant research has found that disorders in signaling pathways can promote cellular autophagy, abnormal osteoclast differentiation, and abnormal cell-cell interactions, leading to oxidative stress and inflammatory responses, etc., which contribute to the formation and progression of osteoporosis [31-33]. In response to the abnormalities in the aforementioned biological processes, we selected the pathways with the top 31 and top 5 FDR values for further in-depth visualization and analysis (Figures 4(B,C)).
Analysis of molecular docking results for core target proteins
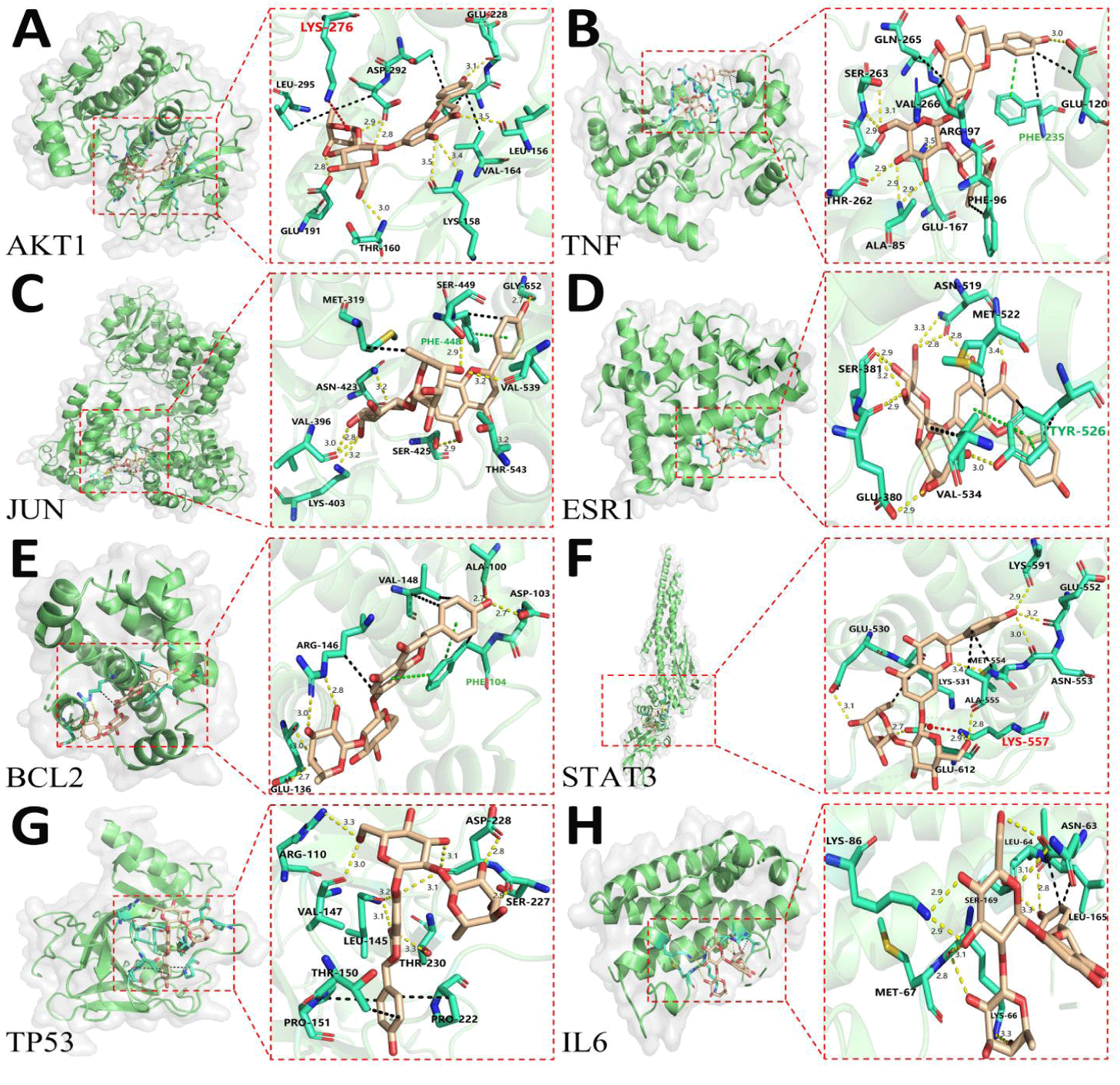
In this study, the core target proteins identified in section 2.2 (TP53, TNF, JUN, STAT3, AKT1, ESR1, BCL2, IL6) underwent virtual screening using AutoDock Vina 1.2.5. The detailed analysis is presented in table 3. Molecular docking analysis revealed significant binding affinity and a high degree of complementarity between NG and the key target proteins (Table 3). Of these, with binding energies all below -7.2 kcal/mol, hydrogen bonds greater than five, and hydrogen bond distances less than 3.6 Å, indicating that the naringin monomer compound has a strong binding effect with a series of core target proteins. The binding energy is a crucial parameter for evaluating the tightness of the interaction between a ligand and a receptor [34]. Generally, the lower the value of the binding energy, the more stable the binding between the two. For these eight core target proteins, their binding energies were all below -7.2 kcal/mol, suggesting very stable binding of NG to these targets. Notably, the binding energies for AKT1 and TNF were particularly low (-9.8 kcal/mol and -9.7 kcal/mol), indicating the tightest binding with NG. The docking scores for AKT1, TNF, BCL2, TP53, and IL6 were superior to those of the original ligand, with scores of -9.8, -9.7, -8.5, -8.0, and -7.3, respectively, suggesting a significant effect of NG on these targets. This confirmed the credibility of the predicted targets for NG treatment of OP through network pharmacology [35]. Subsequently, the PyMOL software was used for visualization, employing yellow dashed lines to represent hydrogen bonds, red dashed lines for salt bridges, and black dashed lines for hydrophobic interactions and the green dashed lines indicate π-π interactions [36].
| Table 3: AutoDock vina results of core target protein. | |||||
| Target | PDB ID | Binding Energy (kcal/mol) | Original Ligand Docking Score | RMSD(Å) | Number of Hydrogen Bonds |
| AKT1 | 4gv1 | -9.8 | -9.4 | 1.178 | 8 |
| TNF | 4y6o | -9.7 | -7.4 | 0.841 | 6 |
| JUN | 6ysv | -9.5 | - | 1.109 | 9 |
| ESR1 | 8du8 | -9.1 | -11.8 | 2.076 | 9 |
| BCL2 | 8c3u | -8.5 | -7.1 | 1.925 | 6 |
| STAT3 | 6NJS | -8.2 | -9.1 | 2.174 | 8 |
| TP53 | 6si0 | -8.0 | -6.2 | 1.817 | 9 |
| IL6 | 1ALU | -7.3 | -5.1 | 1.805 | 9 |
In figure 5A, the results indicated that during simulated docking with the AKT1 receptor, the hydroxyl groups of NG formed eight hydrogen bonds [37] with key residues of the receptor, including ASP-292, GLU-228, LEU-156, LYS-158, THR-160, and GLU-191. Additionally, a stable salt bridge was formed between the LYS-276 residue and the carboxylate group of NG, resulting from electrostatic interactions between the charged residues [38]. The hydrophobic portion of NG also tightly interacted with nonpolar residues GLU-228, ASP-292, LEU-295, and VAL-164 in the hydrophobic pocket of the receptor, promoting the aggregation of hydrophobic molecules [39].
In figure 5B, docking simulations with the TNF receptor showed that the hydroxyl groups of the NG molecule formed six hydrogen bonds with the receptor residues SER-263, THR-262, ALA-85, GLU-167, and GLU-120. The aromatic ring of NG also engaged in Pi-Pi stacking with the aromatic ring of PHE-235 [40], an important non-covalent interaction resulting from the overlap of π-electrons. Furthermore, NG made hydrophobic contacts with residues GLN-265, PHE-235, GLU-120, PHE-96, ARG-97, and VAL-266.
As depicted in figure 5C, docking results revealed that NG, situated in the protein pocket of JUN, formed eight hydrogen bonds with the receptor's VAL-396, LYS-403, SER-425, ASN-423, THR-543, VAL-539, SER-449, and GLY-652 carbonyl groups. Pi-Pi stacking also occurred between the aromatic ring of NG and the aromatic ring of PHE-448. Meanwhile, hydrophobic contacts existed between NG and the MET-319 and SER-449 residues.
As shown in figure 5D, the hydroxyl groups of NG formed nine hydrogen bonds with key residues of the ESR1 receptor, including SER-381, GLU-380, TYR-526, ASN-519, and MET-522. The aromatic residue TYR-526 formed Pi-Pi stacking with the two aromatic rings of NG. Additionally, hydrophobic interactions occurred between NG and MET-522, VAL-534, and TYR-526 residues.
As illustrated in figure 5E, during docking analysis with the BCL2 receptor, the hydroxyl groups of the NG molecule established six hydrogen bonds with the receptor's GLU-136, ARG-146, ALA-100, and ASP-103 key residues, which are crucial for the formation of the stable complex. Meanwhile, the PHE-104 aromatic residue of ESR1 formed Pi-Pi stacking with the aromatic rings of two NG molecules, enhancing the intermolecular binding force through this non-covalent interaction. Moreover, the hydrophobic part of NG also contributed to the stability of the complex through interactions with the nonpolar residues ARG-146, VAL-148, and PHE-104 of the receptor.
In figure 5F, the simulated docking showed that the hydroxyl groups of the NG molecule formed eight hydrogen bonds with the STAT3 receptor's ALA-555, GLU-530, GLU-612, LYS-557, ASN-553, GLU-552, and LYS-591 residues, significantly affecting the binding of the complex. Notably, a stable salt bridge was formed between the LYS-557 residue and carboxylate group of NG. Additionally, the tight interaction between the hydrophobic part of NG and the hydrophobic pocket of the receptor (formed by the nonpolar residues LYS-531, ALA-555, and MET-554) further enhanced intermolecular binding.
In figure 5G, the docking study with the TP53 receptor revealed that the hydroxyl groups of the NG molecule formed nine hydrogen bonds with multiple residues of the receptor (ARG-110, VAL-147, LEU-145, THR-230, ASP-228, and SER-227). The π–π stacking between the PHE-104 aromatic residue and the aromatic rings of the two NG molecules further stabilized the complex structure. Moreover, hydrophobic interactions between NG and the receptor's PRO-151, PRO-222, and THR-150 residues provided additional stability for the complex binding.
In figure 5H, during the docking simulation with the IL6 receptor, the hydroxyl groups of the NG molecule formed nine hydrogen bonds with the LYS-86, MET-67, LYS-66, LEU-165, ASN-63, and SER-169 residues of the receptor, playing an important role in the binding of the complex. The hydrophobic part of the NG molecule also supported the stability of the complex through interactions with the LEU-64 and LEU-165 residues of the receptor.
Discussion
Therapeutic strategies for osteoporosis typically involve pharmacological treatments such as parathyroid hormone and calcitonin [41]. However, these treatments can lead to side effects including varying degrees of renal damage and tissue carcinogenesis [3]. TCM offers a unique perspective and approach to treating OP with fewer side effects [42]. Naringin, a natural flavonoid with a broad range of pharmacological activities, is primarily derived from the peel of Fructus Aurantii and other citrus fruits [4]. Modern research indicated that NG possessed not only antioxidant and anti-inflammatory effects, but also showed potential in combating OP [43]. Its anti-OP mechanisms might affect bone metabolism through various pathways, including promoting osteoblast proliferation and differentiation, inhibiting osteoclast activity, and regulating the expression of factors related to bone metabolism [44-46]. Recently, the combination of active components of TCM and bone tissue engineering has shown promise, with NG/Gelatin Microspheres (GMs)/nano Hydroxyapatite (nHA)/Silk Fibroin (SF) scaffold material emerging as an effective means to promote bone regeneration and repair bone defects [47]. Although NG exhibits extensive activity against OP, the mechanisms of its therapeutic effects were not yet fully understood.
Assisted by network pharmacology, this study used a "Natural Compound-Gene-Target-Disease" model to identify 197 common target genes. Additionally, by employing a "Natural Compound-Core Target" model, we identified eight core targets (TP53, TNF, JUN, STAT3, AKT1, ESR1, BCL2, and IL6). Visualization analysis and virtual docking validation of the core target proteins were conducted to assess whether NG could bind to them. The results revealed that NG demonstrated a high binding affinity for the eight core target genes, suggesting its potential in combating OP. The development and progression of OP are closely related to various inflammatory cytokines [48]. TNF-α, IL-6, and IL-7, among others, have synergistic effects, inducing Receptor Activator of Nuclear Factor-κ B Ligand (RANKL) expression and promoting osteoclast differentiation [49]. TNF-α promotes osteoclast formation and activation by upregulating RANKL binding to RANK on osteoclast precursor cells, thereby increasing bone resorption [50]. Elevated levels of TNF-α and IL-6 during chronic inflammation can lead to a bone metabolic imbalance [51-52]. IL-6 stimulates RANKL production by bone marrow stromal cells, crucial for osteoclast differentiation and activation [53]. It also activates the JAK family, providing binding sites for STAT proteins and affecting estrogen production and function [54]. STAT3, a key intracellular signaling molecule, deletion of the STAT3 gene in bone marrow mesenchymal stem cells or pre-osteoblasts results in craniofacial deformities, osteoporosis, and increased fracture risk in mice [55]. JUN, a member of the AP-1 transcription factor family, plays a significant role in cell proliferation, differentiation, apoptosis, and various signaling pathways. It regulates the RANKL/OPG pathway, modulates osteoclast formation and activity, and interacts with the inflammatory responses, oxidative stress, and hormonal changes [56]. AKT1, BCL2, and TP53 inhibit apoptosis and increase osteoblast survival [57-59]. ESR1 is pivotal in maintaining bone homeostasis through the modulation of multiple signaling pathways [60]; its activation plays a direct role in the physiological processes of osteoblasts and osteoclasts [61], ESR1 helps maintain a dynamic equilibrium between apoptosis and differentiation in these cells. Additionally, it influences the proliferation and differentiation of mesenchymal stem cells into osteoblasts [62]. GO enrichment analysis indicated the involvement of multiple key biological processes in OP development, including signal transduction, apoptosis, cell proliferation, inflammatory responses, and tumor necrosis factor-induced cell responses. Apoptosis, a form of programmed cell death, is crucial for maintaining tissue homeostasis and eliminating damaged cells [63]. NG can promote osteoclast apoptosis through the mitochondrial apoptosis pathway [64] and the regulation of MAPK, ROS, and other signaling pathways [65-66]. Simultaneously, it inhibits osteoblast apoptosis by modulating NF-κB, FOXO, PI3K/Akt, JAK/STAT, Wnt, and other pathways [6-8,67]. Thus, NG may exert its anti-OP effects through the dual regulation of apoptosis via multiple signaling pathways.
KEGG analysis revealed that the pathogenesis of OP is closely linked to abnormalities in various biological processes. Dysfunction of the intracellular signal transduction system triggers a series of chain reactions, including disruptions in autophagy processes, increased oxidative stress levels, intensified inflammatory responses, abnormal osteoclast differentiation, and the disruption of normal communication between bone cells [68]. These factors are not only closely related to the occurrence and development of osteoporosis but are also an indispensable part of its pathological process, involving pathways such as TNF, FoxO, IL-17, PI3K-Akt, AMPK, and VEGF. Activation of the TNF signaling pathway increases osteoclast activity and promotes bone loss. Furthermore, TNF can inhibit osteoblast function, affect bone formation, and accelerate the progression of osteoporosis. Therefore, prolonged activation of the TNF signaling pathway may have negative effects on bone health and increase the risk of osteoporosis [69]. When activated, the FoxO signaling pathway regulates the proliferation and differentiation of bone cells. Inhibition of the FoxO signaling pathway weakens its inhibitory effect on bone cell apoptosis, leading to accelerated bone loss [70]. Activation of the PI3K-Akt signaling pathway promotes osteoblast proliferation and differentiation. It affects cell cycle regulation, apoptosis, and inflammatory responses, thereby indirectly influencing bone metabolism and the development of osteoporosis [71]. The AMPK signaling pathway is closely related to energy and bone metabolism. In vitro studies have shown that AMPK activation promotes osteoblast differentiation and bone matrix mineralization [72]. However, knockout gene studies in mouse models have demonstrated reduced bone density, increased bone formation and resorption, and an overall negative balance of bone remodeling activities, indicating that inhibition of the AMPK signaling pathway may affect the stability of bone tissue [73]. Inhibiting the IL-17 signaling pathway may offer new therapeutic targets for osteoporosis [74]. The VEGF signaling pathway plays a crucial role in angiogenesis by promoting bone repair through the proliferation, migration, and tubule formation of vascular endothelial cells upon VEGF binding to VEGFR [75]. Molecular docking validation showed that the binding between NG and the core target proteins involved hydrogen bonds, salt bridges, π-π stacking, hydrophobic interactions, and other non-covalent interactions. These interactions collectively explain the high binding affinity of naringin for the target proteins. Future experimental studies will further validate these predicted interactions, revealing the molecular mechanisms of NG's action on target protein binding in greater depth.
Conclusion
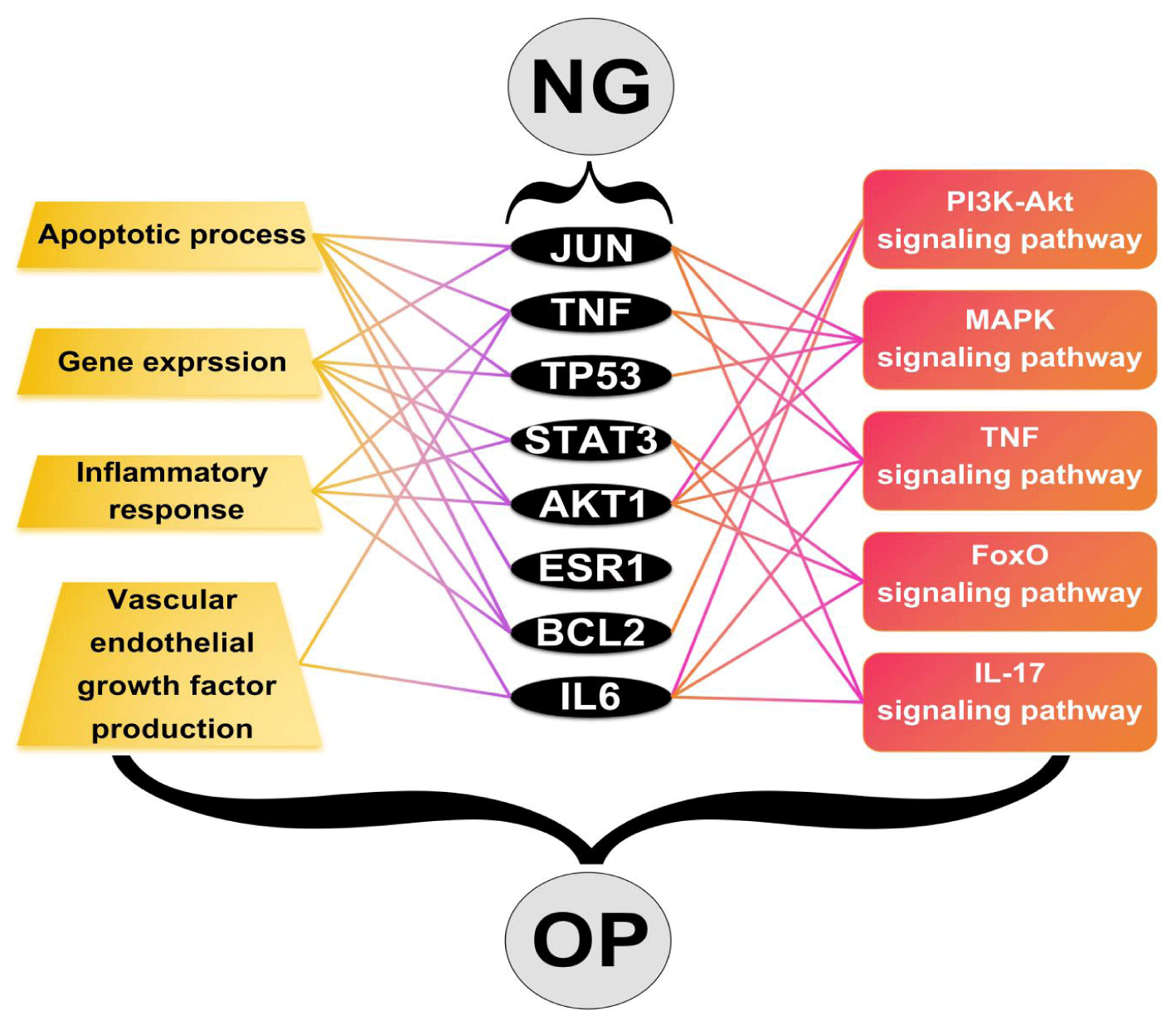
By integrating network pharmacology and molecular docking techniques, this study constructed an analytical framework of "Naringin-Targets-GO and KEGG-Osteoporosis" to predict the potential of Naringin against Osteoporosis. These findings suggest that the underlying molecular mechanisms may involve the regulation of TP53, TNF, JUN, STAT3, AKT1, ESR1, BCL2, and IL6 expression, mediated by signaling pathways such as TNF, FoxO, IL-17, PI3K-Akt, AMPK, and VEGF (Figure 6). These pathways influence multiple biological processes, including the bidirectional regulation of osteoblast proliferation and apoptosis, angiogenesis, and downregulation of inflammation, to achieve anti-OP effects. This implies the pleiotropic effects of NG in modulating bone metabolism and treating OP, offering new directions for treatment and bone tissue engineering. However, there are limitations to this study. First, the database screening process may have overlooked potential NG targets. Second, although GO analysis and KEGG pathway enrichment analysis predicted that NG might affect OP, the specific mechanisms of action require further experimental research for clarification. Third, while this study lays the foundation for future NG research on OP, more work is needed to translate these findings into clinical applications. Future studies will build upon this research, including cell and animal experiments, for validation, aiming for a more accurate understanding. This will facilitate the translation of NG from laboratory research to clinical applications, thereby offering new OP treatment strategies.
Author Contributions
X.L. the design of the study; X.Z. methodology; X.Q., Y.W., W.W. searched the databases, analyzed the data; X.L., X.Y., B.K. writing - original draft preparation; X.L., X.C. graphic design; X.X. resources; ALL. Supervision. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Natural Sciences Foundation of China (Grant No.numbers 82374496). National Traditional Chinese Medicine Experts Inheritance Studio Project (Ministry of Education and Traditional Chinese Medicine [2022] No. 75). Harbin Science and Technology Bureau Project (2023ZCZINS064). Heilongjiang Province Traditional Chinese Medicine Association Young Talent Support Project (2022-QNRC1-12).
Institutional review board statement
Not applicable.
Informed consent statement
Not applicable.
Data availability statement
Data are available from the corresponding authors on reasonable request.
Conflicts of interest
The authors declare no conflicts of interest.
References
- Malisch R, Kotz A. Dioxins and PCBs in feed and food--review from European perspective. Sci Total Environ. 2014 Sep 1;491-492:2-10. doi: 10.1016/j.scitotenv.2014.03.022. Epub 2014 May 5. PMID: 24804623.
- Rivezzi G, Piscitelli P, Scortichini G, Giovannini A, Diletti G, Migliorati G, Ceci R, Rivezzi G, Cirasino L, Carideo P, Black DM, Garzillo C, Giani U. A general model of dioxin contamination in breast milk: results from a study on 94 women from the Caserta and Naples areas in Italy. Int J Environ Res Public Health. 2013 Nov 8;10(11):5953-70. doi: 10.3390/ijerph10115953. PMID: 24217180; PMCID: PMC3863880.
- Omaye ST. Food and nutritional toxicology. Boca Raton: CRC press; 2004.
- Assessment of the health risk of dioxins: Re-evaluation of the Tolerable Daily Intake (TDI). Geneva: World Health Organization. 1998.
- Hoogenboom R, Traag W, Fernandes A, Rose M. European developments following incidents with dioxins and PCBs in the food and feed chain. Food Control. 2015;50:670-683. doi: 10.1016/j.foodcont.2014.10.010.
- Fiedler H. Dioxins in milk, meat, eggs and fish. D'Mello JP, editor. Food Safety: Contaminants and Toxins. Cambridge: CABI Publishing; 2003.
- Lawley R, Curtis L, Davis J. The food safety hazard guidebook second edition. Cambridge: Royal Society of Chemistry Publishing; 2012.
- Schecter A, Cramer P, Boggess K, Stanley J, Päpke O, Olson J, Silver A, Schmitz M. Intake of dioxins and related compounds from food in the U.S. population. J Toxicol Environ Health A. 2001 May 11;63(1):1-18. doi: 10.1080/152873901750128326. PMID: 11346131.
- Parzefall W. Risk assessment of dioxin contamination in human food. Food Chem Toxicol. 2002 Aug;40(8):1185-9. doi: 10.1016/s0278-6915(02)00059-5. PMID: 12067582.
- Szajner J, Czarny-Dzialak M, Dziechciaz M, Pawlas N, Walosik A. Dioxin-Like Compounds (DLCs) in the environment and their impact on human health. Journal of Elementology. 2021;26(2). doi: 10.5601/jelem.2021.26.2.2130.
- Agents classified by the IARC monographs. IARC. 2015
- Alcock RE, Behnisch PA, Jones KC, Hagenmaier H. Dioxin-like PCBs in the environment-human exposure and the significance of sources. Chemosphere. 1998 Oct;37(8):1457-72. doi: 10.1016/s0045-6535(98)00136-2. PMID: 9753761.
- On the implementation of the Community Strategy for dioxins, furans, and polychlorinated biphenyls (COM(2001)593)- Third progress report. Brussel. European Commission. 2010.
- Deshpande S. Handbook of food toxicology. New York: Marcel Dekker Inc; 2002.
- Özbay S, Dikici E, Soylukan C. Evaluation of biological (feed, water), seasonal, and geological factors affecting the heavy metal content of raw milk. Journal of Food Composition and Analysis. 2023;121:105401. doi: 10.1016/j.jfca.2023.105401.
- Exposure to mercury: A major public health concern. Geneva: Public Health and Environment / World Health Organization. 2007.
- Derban LK. Outbreak of food poisoning due to alkyl-mercury fungicide on southern Ghana state farm. Arch Environ Health. 1974 Jan;28(1):49-52. doi: 10.1080/00039896.1974.10666432. PMID: 4586365.
- Järup L. Hazards of heavy metal contamination. British Medical Bulletin. 2003;68:167-182. doi: 10.1093/bmb/ldg032.
- Goyer RA, Clarkson TW. Toxic effects of metals. The McGraw-Hill Companies. 2001.
- Nawrot TS, Staessen JA, Roels HA, Munters E, Cuypers A, Richart T, Ruttens A, Smeets K, Clijsters H, Vangronsveld J. Cadmium exposure in the population: from health risks to strategies of prevention. Biometals. 2010 Oct;23(5):769-82. doi: 10.1007/s10534-010-9343-z. Epub 2010 Jun 3. PMID: 20517707.
- Pan J, Plant JA, Voulvoulis N, Oates CJ, Ihlenfeld C. Cadmium levels in Europe: implications for human health. Environ Geochem Health. 2010 Feb;32(1):1-12. doi: 10.1007/s10653-009-9273-2. Epub 2009 Aug 18. PMID: 19688602.
- Exposure of cadmium: A major public health. Geneva: Public Health and Environment World Health Organization. 2010.
- Amundsen PA, Staldvik FJ, Lukin AA, Kashulin NA, Popova OA, Reshetnikov YS. Heavy metal contamination in freshwater fish from the border region between Norway and Russia. Sci Total Environ. 1997 Aug 18;201(3):211-24. doi: 10.1016/s0048-9697(97)84058-2. PMID: 9241871.
- Yılmaz F. The comparison of heavy metal concentrations (Cd, Cu, Mn, Pb, and Zn) in tissues of three economically important fish (anguilla anguilla, mugil cephalus and oreochromis niloticus) inhabiting köycegiz lake-mugla (Turkey). Turkish Journal of Science & Technology. 2009;4(1):7-15.
- Groten JP. Adverse effect of food contaminants. Vries JD. Editor. Food Safety and Toxicity. Boca Raton: CRC Press LLC; 1997.
- Oymak T, Tokalıoğlu Ş, Yılmaz V, Kartal Ş, Aydın D. Determination of lead and cadmium in food samples by the coprecipitation method. Food Chemistry. 2009;113:1314-1317. doi: 10.1016/j.foodchem.2008.08.064.
- Thompson LJ, Gupta içinde RC. Veterinary toxicology. 2012:522-526. doi: 10.1016/C2010-0-67763-7.
- Demirci M, Gıda Kimyası. Tekirdağ: Rebel yayıncılık. 2003.
- Chi QQ, Zhu GW, Alan L. Bioaccumulation of heavy metals in fishes from Taihu Lake, China. J Environ Sci (China). 2007;19(12):1500-4. doi: 10.1016/s1001-0742(07)60244-7. PMID: 18277656.
- Zhuang P, Zou H, Shu W. Biotransfer of heavy metals along a soil-plant-insect-chicken food chain: field study. J Environ Sci (China). 2009;21(6):849-53. doi: 10.1016/s1001-0742(08)62351-7. PMID: 19803093.
- Cramér K, Goyer RA, Jagenburg R, Wilson MH. Renal ultrastructure, renal function, and parameters of lead toxicity in workers with different periods of lead exposure. Br J Ind Med. 1974 Apr;31(2):113-27. doi: 10.1136/oem.31.2.113. PMID: 4830763; PMCID: PMC1009566.
- Janssen MM. Contaminantlar. Vries JD, editor. Foof Safety and Toxicity. Boca Raton: CRC Press LLC; 1997.
- Anawar HM, Akai J, Mostofa KM, Safiullah S, Tareq SM. Arsenic poisoning in groundwater: health risk and geochemical sources in Bangladesh. Environ Int. 2002 Feb;27(7):597-604. doi: 10.1016/s0160-4120(01)00116-7. PMID: 11871394.
- Erol İ. Gıda hijyeni ve mikrobiyolojisi. Ankara: Pozitif Matbaacılık (In Turkish). 2007.
- Aposhian HV, Aposhian MM. Newer developments in arsenic toxicity. Journal Of The Amerıcan College Of Toxıcology. 1989;8(7):1297-1305.
- Goering PL, Aposhian HV, Mass MJ, Cebrián M, Beck BD, Waalkes MP. The enigma of arsenic carcinogenesis: role of metabolism. Toxicol Sci. 1999 May;49(1):5-14. doi: 10.1093/toxsci/49.1.5. PMID: 10367337.
- Roychowdhury T, Uchino T, Tokunaga H, Ando M. Survey of arsenic in food composites from an arsenic-affected area of West Bengal, India. Food Chem Toxicol. 2002 Nov;40(11):1611-21. doi: 10.1016/s0278-6915(02)00104-7. PMID: 12176088.
- Duxbury JM, Mayer AB, Lauren JG, Hassan N. Food chain aspects of arsenic contamination in Bangladesh: effects on quality and productivity of rice. J Environ Sci Health A Tox Hazard Subst Environ Eng. 2003 Jan;38(1):61-9. doi: 10.1081/ese-120016881. PMID: 12635819.
- Ng JC, Wang J, Shraim A. A global health problem caused by arsenic from natural sources. Chemosphere. 2003 Sep;52(9):1353-9. doi: 10.1016/S0045-6535(03)00470-3. PMID: 12867164.
- Ratnaike RN. Acute and chronic arsenic toxicity. Postgrad Med J. 2003 Jul;79(933):391-6. doi: 10.1136/pmj.79.933.391. PMID: 12897217; PMCID: PMC1742758.
- Saha JC, Dikshit AK, Bandyopadhyay M, Saha KC. A review of arsenic poisoning and its effects on human healt. 1999;29(3):281-313. doi: 10.1080/10643389991259227.
- Yoshida T, Yamauchi H, Fan Sun G. Chronic health effects in people exposed to arsenic via the drinking water: dose-response relationships in review. Toxicol Appl Pharmacol. 2004 Aug 1;198(3):243-52. doi: 10.1016/j.taap.2003.10.022. PMID: 15276403.
- Cverenkárová K, Valachovičová M, Mackuľak T, Žemlička L, Bírošová L. Microplastics in the Food Chain. Life (Basel). 2021 Dec 6;11(12):1349. doi: 10.3390/life11121349. PMID: 34947879; PMCID: PMC8704590.
- van Raamsdonk LWD, van der Zande M, Koelmans AA, Hoogenboom RLAP, Peters RJB, Groot MJ, Peijnenburg AACM, Weesepoel YJA. Current Insights into Monitoring, Bioaccumulation, and Potential Health Effects of Microplastics Present in the Food Chain. Foods. 2020 Jan 9;9(1):72. doi: 10.3390/foods9010072. PMID: 31936455; PMCID: PMC7022559.
- Barboza LGA, Vieira LR, Branco V, Figueiredo N, Carvalho F, Carvalho C, Guilhermino L. Microplastics cause neurotoxicity, oxidative damage and energy-related changes and interact with the bioaccumulation of mercury in the European seabass, Dicentrarchus labrax (Linnaeus, 1758). Aquat Toxicol. 2018 Feb;195:49-57. doi: 10.1016/j.aquatox.2017.12.008. Epub 2017 Dec 20. PMID: 29287173.
- Pironti C, Ricciardi M, Motta O, Miele Y, Proto A, Montano L. Microplastics in the Environment: Intake through the Food Web, Human Exposure and Toxicological Effects. Toxics. 2021 Sep 16;9(9):224. doi: 10.3390/toxics9090224. PMID: 34564375; PMCID: PMC8473407.
- Prata JC, Reis V, da Costa JP, Mouneyrac C, Duarte AC, Rocha-Santos T. Contamination issues as a challenge in quality control and quality assurance in microplastics analytics. J Hazard Mater. 2021 Feb 5;403:123660. doi: 10.1016/j.jhazmat.2020.123660. Epub 2020 Aug 9. PMID: 33264868.
- Walkinshaw C, Lindeque P, Thompson R, Tolhurst T, Cole M. Microplastics and seafood: Lower trophic organisms at highest risk of contamination. Ecotoxicology and environmental safety. 2019;190:110066. doi: 10.1016/j.ecoenv.2019.110066.
- Habib RZ, Kindi RA, Salem FA, Kittaneh WF, Poulose V, Iftikhar SH, Mourad AI, Thiemann T. Microplastic Contamination of Chicken Meat and Fish through Plastic Cutting Boards. Int J Environ Res Public Health. 2022 Oct 18;19(20):13442. doi: 10.3390/ijerph192013442. PMID: 36294029; PMCID: PMC9602623.
- Pei X, Tandon A, Alldrick A, Giorgi L, Huang W, Yang R. The China melamine milk scandal and its implications for food safety regulation. FoodPolicy. 2011;36(3):412-420. doi: 10.1016/j.foodpol.2011.03.008.
- Ingelfinger JR. Melamine and the global implications of food contamination. N Engl J Med. 2008 Dec 25;359(26):2745-8. doi: 10.1056/NEJMp0808410. PMID: 19109571.
- Newton GL, Utley PR. Melamine as a dietary nitrogen source for ruminants. Journal of Animal Science. 1978;47(6):1338-1344. doi: 10.2527/jas1978.4761338x. Pei X, Tandon A, Alldrick A, Giorgi L, Huang W, Yang R. The China melamine milk scandal and its implications for food safety regulation. Food Policy. 2011;36(3):412-420. doi: 10.1016/j.foodpol.2011.03.008.
- Evans JR, Clarke VC. The nitrogen cost of photosynthesis. J Exp Bot. 2019 Jan 1;70(1):7-15. doi: 10.1093/jxb/ery366. PMID: 30357381.
- Shukla S, Saxena A. Sources and leaching of nitrate contamination in groundwater. Current Science. 2020;118(6):883-891. doi: 10.18520/cs/v118/i6/883-891.
- Horsch AM, Sebranek JG, Dickson JS, Niebuhr SE, Larson EM, Lavieri NA, Ruther BL, Wilson LA. The effect of pH and nitrite concentration on the antimicrobial impact of celery juice concentrate compared with conventional sodium nitrite on Listeria monocytogenes. Meat Sci. 2014 Jan;96(1):400-7. doi: 10.1016/j.meatsci.2013.07.036. Epub 2013 Aug 3. PMID: 23973624.
- Hill MJ. Nitrates and nitrites in food and water (Vol. 7). CRC Press; 1996.
- Walker R. The metabolism of dietary nitrites and nitrates. Biochem Soc Trans. 1996 Aug;24(3):780-5. doi: 10.1042/bst0240780. PMID: 8878847.
- Andrade R, Reyes FG, Rath S. A method for the determination of volatile N-nitrosamines in food by HS-SPME-GC-TEA. Food Chemistry. 2005;91(1):173-179. doi: 10.1016/j.foodchem.2004.08.015.
- Archer DL. Evidence that ingested nitrate and nitrite are beneficial to health. J Food Prot. 2002 May;65(5):872-5. doi: 10.4315/0362-028x-65.5.872. PMID: 12030305.
- Honikel KO. The use and control of nitrate and nitrite for the processing of meat products. Meat Sci. 2008 Jan;78(1-2):68-76. doi: 10.1016/j.meatsci.2007.05.030. Epub 2007 Jun 27. PMID: 22062097.
- Lundberg JO, Larsen FJ, Weitzberg E. Supplementation with nitrate and nitrite salts in exercise: a word of caution. J Appl Physiol (1985). 2011 Aug;111(2):616-7. doi: 10.1152/japplphysiol.00521.2011. PMID: 21828255.
- Özbay S. Investigation of volatile N-nitrosamine contents of hams with different ingredients produced in Turkey. Journal of Food Composition and Analysis. 2024;130:106170. doi: 10.1016/j.jfca.2024.106170
- Renner R. Perchlorate in food. Environ Sci Technol. 2008 Mar 15;42(6):1817. doi: 10.1021/es0870552. PMID: 18409597.
- Dasgupta PK, Martinelango PK, Jackson WA, Anderson TA, Tian K, Tock RW, Rajagopalan S. The origin of naturally occurring perchlorate: the role of atmospheric processes. Environ Sci Technol. 2005 Mar 15;39(6):1569-75. doi: 10.1021/es048612x. PMID: 15819211.
- Blount BC, Valentin-Blasini L, Osterloh JD, Mauldin JP, Pirkle JL. Perchlorate exposure of the US Population, 2001-2002. J Expo Sci Environ Epidemiol. 2007 Jul;17(4):400-7. doi: 10.1038/sj.jes.7500535. Epub 2006 Oct 18. PMID: 17051137.
- Wolff J. Perchlorate and the thyroid gland. Pharmacol Rev. 1998 Mar;50(1):89-105. PMID: 9549759.
- Charnley G. Perchlorate: overview of risks and regulation. Food Chem Toxicol. 2008 Jul;46(7):2307-15. doi: 10.1016/j.fct.2008.03.006. Epub 2008 Mar 10. PMID: 18440116.
- Greer MA, Goodman G, Pleus RC, Greer SE. Health effects assessment for environmental perchlorate contamination: the dose response for inhibition of thyroidal radioiodine uptake in humans. Environ Health Perspect. 2002 Sep;110(9):927-37. doi: 10.1289/ehp.02110927. Erratum in: Environ Health Perspect. 2005 Nov;113(11):A732. PMID: 12204829; PMCID: PMC1240994.
Content Alerts
SignUp to our
Content alerts.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.